I.INTRODUCTION
Epilepsy is a progressive neurological condition that millions of people live with, as it is associated with repeated occurrences of unexpected seizures. Electroencephalogram (EEG) is still the best solution in diagnosing and monitoring epilepsy because it is non-invasive and it shows the electrical activity of the brain. The analysis of EEG signals to detect seizures can help automate the identification process, which would otherwise require diagnostic delay and attention from neurologists, and it would also allow for maintaining continuous working systems. Over the last several years, Machine Learning (ML) and DL have held great potential in this area, and many studies have investigated their performance at EEG signal classification in a variety of neurological conditions [1].
Nonetheless, the research and development of a clinically viable automated seizure detection system presents a 2-fold problem, that of finding a highly accurate system and a highly interpretable system. Although deep learning techniques such as convolutional neural networks (CNNs) and transformers can achieve high accuracy (close to 100 percent) on raw EEG time-series data, they operate as black box/non-interpretable models. Such opacity is one of the greatest impediments to clinical application, since doctors need evidence they can comprehend in order to be willing to take a model at its prediction. In addition, the presence of severe methodological concerns in the assessment of the model, like the insufficient use of proper cross-validation procedures, may cause overly optimistic performance estimates that would not necessarily translate to new patients, which is crucial to the process of translation of the technology into the real world [2].
Conversely, in traditional ML pipelines, global feature selection methods, such as principal component analysis (PCA), or sequential feature selection (SFS) have been used. The methods minimize the dimensionality, but they do not take into consideration the neurophysiological topology of the brain. They are also processing all EEG channels as an amorphous group of inputs, possibly overlooking clinically useful signals of clinically essential regions, for example, the temporal lobe in focal epilepsy. Moreover, the majority of the existing models, particularly, the ones developed to assess a cross-patient application, fail to incorporate valuable demographic and clinical metadata of the patients (e.g., age, gender, and medication history), which could substantially affect EEG data patterns and enhance the generalization properties of the model, an issue that patient-adversarial networks have started to target [3].
The proposed research pertains to the need of a seizure detection tool that is accurate, interpretable, and possesses a spectrum of clinical awareness. It suggests a newly developed framework that explicitly maintains the neurophysiological setting of the EEG recordings with the metadata of the patient. We mainly have four contributions to make. We propose the region-aware feature selection (RAFS) scheme based on a genetic algorithm (GA) that acts locally, on a brain part-by-part, leading to the fact that the resulting feature set is minimal in size, yet clinically interpretable. Age, gender, medication, and seizure history data of the patient can also be complemented with our model, incorporating patient metadata in a systematic way with the chosen EEG features to reflect the overall picture of the patient. Moreover, an ensemble of robust classifier models, with RandomForest, ExtraTrees, and XGBoost, with high performance, is used to attain a top 1 accuracy on the SNP-wise basis-broken-down data. This study proposes a paradigm that balances between the predictive performance and the relative interpretability, leading to a solution that is realistic to adopt and honest in its reliability regarding autonomous EEG seizure detection.
A.PROBLEM OF EXISTING METHODS
The existing automated seizure detectors introduce a major trade-off between their performance and clinical applicability. There are, on the one hand, deep learning models such as CNNs and transformers, which are highly accurate, yet work as black boxes, thus consuming a great deal of computing resources, such as Graphics Processing Unit (GPUs) to manipulate raw time-series data, which prevents their transparency and demonstrability in practice. Conversely, machine learning pipelines using global feature selection (based on PCA or SFS, and so on) are commonly seen in traditional applications and, although more interpretable (i.e., readily identifiable as original features), all EEG channels are treated as an undifferentiated mass. There is a problem in misrepresenting the neurophysiological surface of the brain, underlying the neurophysiology and may lose signals of areas of clinical importance. Moreover, most of the available models, regardless of paradigm, do not incorporate useful patient-specific metadata such as age or gender, or medication factors, and as such, they have limited capacities to generalize to different populations of patients and real clinical settings.
B.NOVELTY
This study proposes a region-sensitive GA-inspired feature selection algorithm of EEG seizure detection in which region-wise superior EEG channels are incorporated with the demographic metadata of the EEG data. When integrated with a lightweight soft voting ensemble, the proposed method results in a 99.28% accuracy on the images of the chest with these compact yet physiologically relevant features, surpassing traditional ML-based algorithms and reducing the accuracy of the deep CNN counterpart to the same level with only a fraction of the training and inference time, finally allowing the proposed method to be deployed in a clinical setting.
II.RELATED WORKS
The automated seizure detection has developed rather fast with the shift in the study focus to complex AI-powered detection techniques. A large literature has dealt with the idea of integrating the feature-extracting potential of deep learning with the efficiency of the classical classifiers. As an example, hybrid CNN or Deep Neural Network (DNN) feature extractor models can be followed by a support vector machine (SVM) with or without dimensionality reduction in the form of PCA, which have shown a high level of accuracy [4]. There are still other automated systems, which apply many forms of machine learning algorithms to the processed EEG data in order to be able to detect aberrant epileptic episodes effectively [5].
One of the main drawbacks of a significant number of high-performing deep learning models is that they are not interpretable. In order to mitigate this detrimental gap, scientists have started turning to explainable AI (XAI) practices. Better visualization of the spectrogram interpretability of the patient-independent seizure classification of deep learning models has also been created [6], and more advanced algorithms in XAI provide a spectrum and spatio-temporal interpretation of deep learning decisions [7]. This has been a growing trend and has been highlighted in comprehensive reviews [8], and the deep learning-based seizure detection methodology remains an active research field [9].
Deep neural networks have proven to be especially successful in targeted scenarios, including the review of 2-channel EEG to identify the occurrence of a seizure, indicating just how feasible they will be in low-resource or home settings [10]. Additional advanced architectures are being explored as well to address bespoke problems, for example, long-term EEG partitioning networks are proposed to sharpen the accuracy of detecting seizure onset by considering long durations of recording [11]. Along with the strategy of coupling CNN-based deep feature extraction and shallow, efficient classifiers, this approach has been a popular and effective one in the literature [12]. There is also a shift toward predicting instead of just detecting, with deep learning models being created in order to predict neonatal seizures based on their EEGs at a short horizon [13]. In order to further increase the reliability of the model, one of the ways to state the uncertainty in seizure detection based on EEG was suggested, called evidential multi-view learning, which is also aimed at quantifying the uncertainty [14].
The architectural landscape of the models is getting more diverse. There is also considerable promise in multi-representation deep learning approaches that consume multi-format input to achieve a more diverse set of features [15]. One of the most important steps in moving toward the creation of robust and reliable models is having in place pipelines by which models can be retrained and re-evaluated consistently on new data [16]. The influence of deep learning skills and accessibility of data has been reviewed in an orderly manner as well [17]. Some studies have also discussed the application of different EEG modalities, where machine learning and deep learning methods have also been applied to detecting seizures due to intracranial electroencephalography (iEEG) [18].
The aspect of hybrid models that integrates the advantages of various regimes is a very dynamic research field. A case in point, hybrid architectures that combine XGBoost and recurrent neural networks (RNNs) are under consideration to increase the performance of seizure detection [19]. In the same line, more complex and accurate recognition of seizures with the use of the EEG signal is evolving toward hybrid CNN-BiLSTM models [20]. Transformer-based frameworks are also moving into EEG analysis, where models such as CNN-Informer on long-term recordings [33] and transformer-based wavelet-scalograms are on the cusp of excellent seizure recognition scores [21]. Directly addressing the problem of interpretability, channel-annotated deep learning models are being developed in order to explicitly connect the predictions to the input of individual EEG channels [22].
EEG analysis is expanding as well in its range to cover other linked neuronal conditions. One application that relies on hybrid algorithms of deep learning is the early seizure detection caused by stress and anxiety, where the input of the EEG with position data is enhanced [23]. The detection of seizures is also being enhanced by multi-input deep feature learning networks [24]. Federated machine learning is under research for this reason [25]. Highly annotated, high-quality data are the key to solving the problem, which is also emphasized in comprehensive reviews of the published EEG datasets [26]. In addition to seizure detection, measures of functional connectivity are being used to predict the onset of zones of seizures based on interictal iEEG [27].
One of the primary aspects of utilizing these models in practice is their technology, which makes it possible to develop smart Internet of Things (IoT)-cloud frameworks with adaptive deep learning [28]. The huge amount of research has been summarized in systematic reviews and meta-analyses [29]. SVMs are being integrated with specific deep learning architectures such as EfficientNet-B0 [30], and the possibility of making an impact on the real world is being fulfilled in the form of systems that achieve this goal with SVMs and IoT-wearable EEG devices [31]. Comparison of automated machine learning (AutoML) tools has also been done in order to facilitate the streamlining of the development of the model [32].
Lastly, the innovation will be provided by adding dual attention mechanisms to concentrate on salient features [34] as well as testing other data representations in the context of this task, for example, an imaging form of an EEG signal [35]. The range of methods is considerable, with everything including what has been named physiological signals [36], sentiment analysis [36], and the more exotic concept of signal processing and strange attractors through deep learning [39]. Design of new feature extraction entries into the machine learning models is a high-priority research area [37], with others even considering cross-species and cross-modality schemes [38]. Finally, the research activity on both machine learning and deep learning methods is still being developed along the same line of automated detection of epileptic seizures based on EEG signals [40].
The given proposal is situated in the synthesis of these tendencies. It accounts for the drawbacks of the current strategies deployed to include a syllabic method of region-to-region feature selection that survives fundamentally diverse to global algorithms, such as PCA or purely data-driven deep learning techniques. Our framework offers a highly accurate, clinically interpretable, and computationally efficient solution that can fill the gap between the most well-known black-box structures of deep learning and more traditional and less accurate pipelines of machine learning, considering both demographic metadata and a lightweight and versatile combination of algorithms.
III.MATERIALS AND METHODS
This section provides a detailed description of the dataset, preprocessing steps, and the overall methodological framework employed in this study.
A.MATERIALS
1).DATASET DESCRIPTION
This study will be carried out on a publicly available study on EEG recordings and demographic metadata of 50 drug-resistant epilepsy sufferers, which can be found on the IEEE Dataport site (“Early Warning of Epileptic Seizure Onset”). The specified dataset can be of great value due to the age range of patients that is large (1–90 years) and the crucial clinical metadata, usually unavailable in benchmark datasets such as Bonn or Children’s Hospital Boston—Massachusetts Institute of Technology (CHB-MIT). Informed consent was utilized in the collection of all the data, and no patient identifiers were given, so the data are anonymous.
The initial dataset is made up of 100,000 samples, with each one being the representation of a snapshot of EEG activity. All the samples are described by 27 features. Among these are 22 features describing the amplitude of EEG channels (the difference between the electrical potentials on two electrodes: e.g., FP1-F7, C3-P3, and T8-P8-1). Each sample possesses four demographic and clinical meta-features: age, gender, medication status, and seizure history, besides the EEG data that it comprises. The last binary label identifies the state of the sample as being a non-seizure (0) or seizure (1). In our region-aware processing, the 22 EEG channels were partitioned into four particular neurophysiological regions according to the standard brain topology. Our feature selection strategy requires this grouping, and it is outlined in Table I.
Table I. Neurophysiological grouping of EEG channels
| Brain region | EEG channels included |
|---|---|
| Frontal | ‘FP1-F7’, ‘FP1-F3’, ‘FP2-F4’, ‘FP2-F8’, ‘F3-C3’, ‘F4-C4’, ‘FZ-CZ’ |
| Central-parietal | ‘C3-P3’, ‘C4-P4’, ‘CZ-PZ’, ‘P3-O1’, ‘P4-O2’ |
| Temporal | ‘F7-T7’, ‘F8-T8’, ‘FT10-T8’, ‘FT9-FT10’, ‘T7-FT9’, ‘T7-P7’, ‘T8-P8-0’, ‘T8-P8-1’ |
| Occipital | ‘P7-O1’, ‘P8-O2’ |
2).DATA PREPROCESSING AND AUGMENTATION
All EEG features were standardized using Z-score normalization to ensure comparable feature scales and stable model training. The equation of Z-score normalization is the following:
where x is the original feature value, mu is the mean of the feature column, and sigma is the standard deviation of the feature column. No data augmentation was performed, as the study focuses on feature selection from the original processed amplitudes.B.METHODOLOGICAL FRAMEWORK
Experiments were done in a Python-based environment with the usage of the scikit-learn library to implement the machine learning models and the DEAP library to implement the GA. This combination of well-established libraries will offer a strong and repeatable system to construct and analyze our desired pipeline. All the course of the work, including data loading and model evaluation, was scripted so that the outcome would coincide with all the trial runs.
IV.PROPOSED METHODOLOGY
In this part, we describe the architecture of our framework that detects EEG seizures. The end-to-end pipeline that the system was designed for converts raw EEG and demographic data into the correct and comprehensible classification, as shown in Fig. 1.
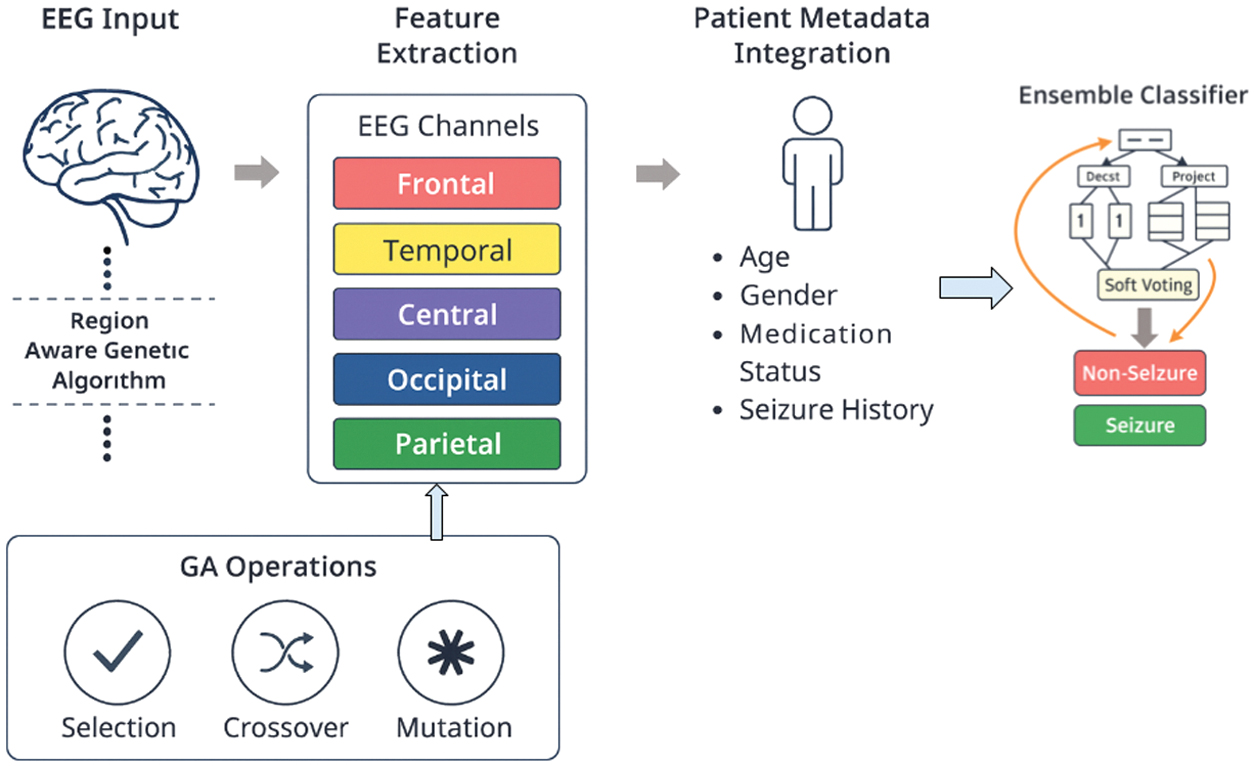 Fig. 1. Overview of the proposed region-aware GA-based EEG seizure detection framework with feature selection, meta-feature integration, and ensemble classification.
Fig. 1. Overview of the proposed region-aware GA-based EEG seizure detection framework with feature selection, meta-feature integration, and ensemble classification.
A.SYSTEM MODEL
The new system suggested is a multi-stage diagnostic pipeline that is expected to put a lot of emphasis on clinical relevance as well as predictive accuracy. Within Brain Region Segregation, the 22 channels of EEG data are assigned to the four preselected neurophysiological regions that are preselected. The second step is the actual innovation of our system; that is, region-aware GA feature selection. In such a configuration, a corresponding GA is run on each region of the brain to determine the two most effective channels, and it is again left with eight optimal EEG characteristics. Then, during the Meta-Feature Integration process, the eight of these channels are integrated with the four demographic and clinical meta-features to produce a final, smaller feature 12-feature vector. The last phase is Ensemble Learning and Classification, which may apply this 12-feature dataset to train a soft-voting type of ensemble classifier, which consists of RandomForest, ExtraTrees, and XGBoost, in order to produce the final robust and accurate result of classification.
B.GA FOR REGION-WISE FEATURE SELECTION
GA is a heuristic search algorithm that is based on the theory of natural evolution by Charles Darwin. It is also adapted quite well to complicated optimization problems such as feature selection. Our solution is the first of its type applied on a region-by-region basis. And, in both brain regions, the population, under the GA, will be composed of people (chromosomes) each of which is a binary vector and whose components are bound to choosing two and only two of the channels. The fitness of a particular chromosome is measured by the accuracy of classification of a basic RandomForest classifier trained on only the channels chosen in that chromosome. The population is then refined by the GA in repeated evolutionary operations. The chromosomes with the better fitness are more likely to be reproduced (selection), the pair of parent chromosomes exchange material (crossover) to produce the offspring, and some error in the chromosomes will be randomly introduced (mutation), to keep some diversity and cover the search space. This evolution undergoes evolution as explained in Algorithm 1 until the algorithm stops at the optimal set of two channels in each region.
GA hyperparameters were set to POP = 15, GEN = 6, CXPB = 0.5, and MUTPB = 0.2. Fitness was evaluated as mean 3-fold CV accuracy of a RandomForest classifier, and the algorithm currently terminates after a fixed number of generations (GEN = 6):
Algorithm 1. Region-Wise Genetic Algorithm for Feature Selection
| • : EEG data for a single region |
| • : Population size |
| • : Number of generations |
| • : Crossover probability |
| • : Mutation probability |
| • : A set of 2 optimal channels for the region. |
| 1. |
| 2. Evaluate Fitness: for each chromosome in P do: |
| features ← Select channels from Data_region based on chromosome. |
| fitness_score ← Train and evaluate a baseline classifier (e.g., RandomForest) using features.chromosome. |
| fitness ← fitness_score. |
| end for |
| 3. Evolve for Generations: |
| for g from 1 to N_gen do: |
| a. Selection: Select N_pop individuals from P to form a mating pool, M, based on their fitness scores (e.g., tournament selection). |
| b. Crossover: |
| for each pair of parent1, parent2 in M do: |
| if random() < CX_PB then: |
| child1, child2 ← Perform crossover on parent1, parent2. |
| Add child1, child2 to a new population P_new. |
| end if |
| end for |
| c. Replacement: |
| Replace the old population P with P_new. |
| d. Evaluate Fitness: |
| Recalculate fitness for all individuals in the new population P. |
| end for |
| 4. Return Best Individual: best_individual ← Select the individual with the highest fitness from the final population P. |
| best_channels ← Decode to get the final channel set. |
| return . |
C.ENSEMBLE LEARNING
Ensemble learning is a way in which several machine learning models predict together to give a better ground prediction by averaging. The effectiveness and simplicity of soft-voting ensembles prompted us to use it as a composition of three tree-based, strong and diverse models. The first, RandomForest, is, again, the ensemble approach that works by building a vast number of decision trees during the training time. The output of the RandomForest of a classification task is the class that is selected by the majority of trees. It is a correction of the tendency of decision trees to overfit to the training set. The second, ExtraTrees (Extremely Randomized Trees), is more or less like RandomForest, but it adds increased randomness in the choice of node split points. It chooses a random split at each feature instead of calculating the locally optimal one, and, therefore, it may decrease variance and increase the predictive accuracy. The third, XGBoost (Extreme Gradient Boosting) is an extremely efficient and powerful realization of the gradient boosting framework. Its method is to create each model sequentially and train the new one to produce a correction to the previous models, thus forming a very powerful composite model.
A soft-voting ensemble gives the probability of every class, given by each individual classifier. These probabilities are averaged, and the one with the largest average is the final prediction, as done in Algorithm 2. This scheme uses the soft scores in the form of confidence of each individual model and can outperform hard voting in many cases:
Algorithm 2. Soft-Voting Ensemble Classifier
| Input: |
| • : A single data point with 12 features |
| • : Trained RandomForest model |
| • : Trained ExtraTrees model |
| • : Trained XGBoost model |
| • : The predicted class label (‘Seizure’ or ‘Non-Seizure’). |
| 1. Get Individual Probabilities: |
| Prob_RF ← M_RF.predict_proba(test_sample) |
| Prob_ET ← M_ET.predict_proba(test_sample) |
| Prob_XGB ← M_XGB.predict_proba(test_sample) |
| 2. Average the Probabilities: |
| Avg_Prob_NonSeizure ← ([0] + [0] + [0]) / 3 |
| Avg_Prob_Seizure ← (Prob_RF[1] + Prob_ET[1] + Prob_XGB[1]) / 3 |
| 3. Make Final Prediction: |
| if Avg_Prob_Seizure > Avg_Prob_NonSeizure then: |
| final_prediction ← ‘Seizure’ |
| else: |
| final_prediction ← ‘Non-Seizure’ |
| end if |
| 4. return . |
V.EXPERIMENTAL RESULTS
The reported section represents the performance assessment of our suggested framework. We describe the preparation of experiments, quantitative parameters, and a comparison of individual models to the final ensemble.
A.ENVIRONMENTAL SETUP AND HYPERPARAMETER TUNING
The dataset was partitioned using a 70/10/20 train–validation–test split, and all models were further evaluated using 5-fold cross-validation. The performance variation across folds was within ± 0.1 to 0.37 %%, indicating high stability and robustness of the proposed framework. Ensemble models were tuned with a grid search and cross-validation on the hyperparameters of the models. The hyperparameters listed in Table II are the most significant ones.
Table II. Hyperparameter configuration for ensemble models
| Model | Key hyperparameter | Value |
|---|---|---|
| RandomForest | n_estimators | 100 |
| max_depth | None | |
| ExtraTrees | n_estimators | 100 |
| max_depth | None | |
| XGBoost | n_estimators | 100 |
| learning_rate | 0.1 | |
| max_depth | 5 | |
| For all models | Random seed | 42 |
| Cross-validation | 5 |
B.PERFORMANCE EVALUATION
The performance of the individual classifiers and the final soft-voting ensemble was evaluated using accuracy as the primary metric. The results, shown in Table III, demonstrate the superiority of the ensemble approach. Its graphical representation is shown in Fig. 2.
| Class | Precision (mean ± Std) | Recall (mean ± Std) | F1-Score (mean ± Std) | Accuracy (mean ± Std) | Model |
|---|---|---|---|---|---|
| 0 | 98.78% | 99.81% | 99.29% | 99.29 ± 0.1% | Voting Ensemble |
| 1 | 99.81% | 98.77% | 99.29% | ||
| 0 | 98.93% | 99.03% | 98.98% | 98.97 ± 0.25% | RandomForest |
| 1 | 99.01% | 98.91% | 98.96% | ||
| 0 | 98.64% | 98.82% | 98.73% | 98.72 ± 0.32% | ExtraTrees |
| 1 | 98.80% | 98.62% | 98.71% | ||
| 0 | 98.34% | 98.50% | 98.42% | 98.41 ± 0.37% | XGBoost |
| 1 | 98.48% | 98.32% | 98.40% |
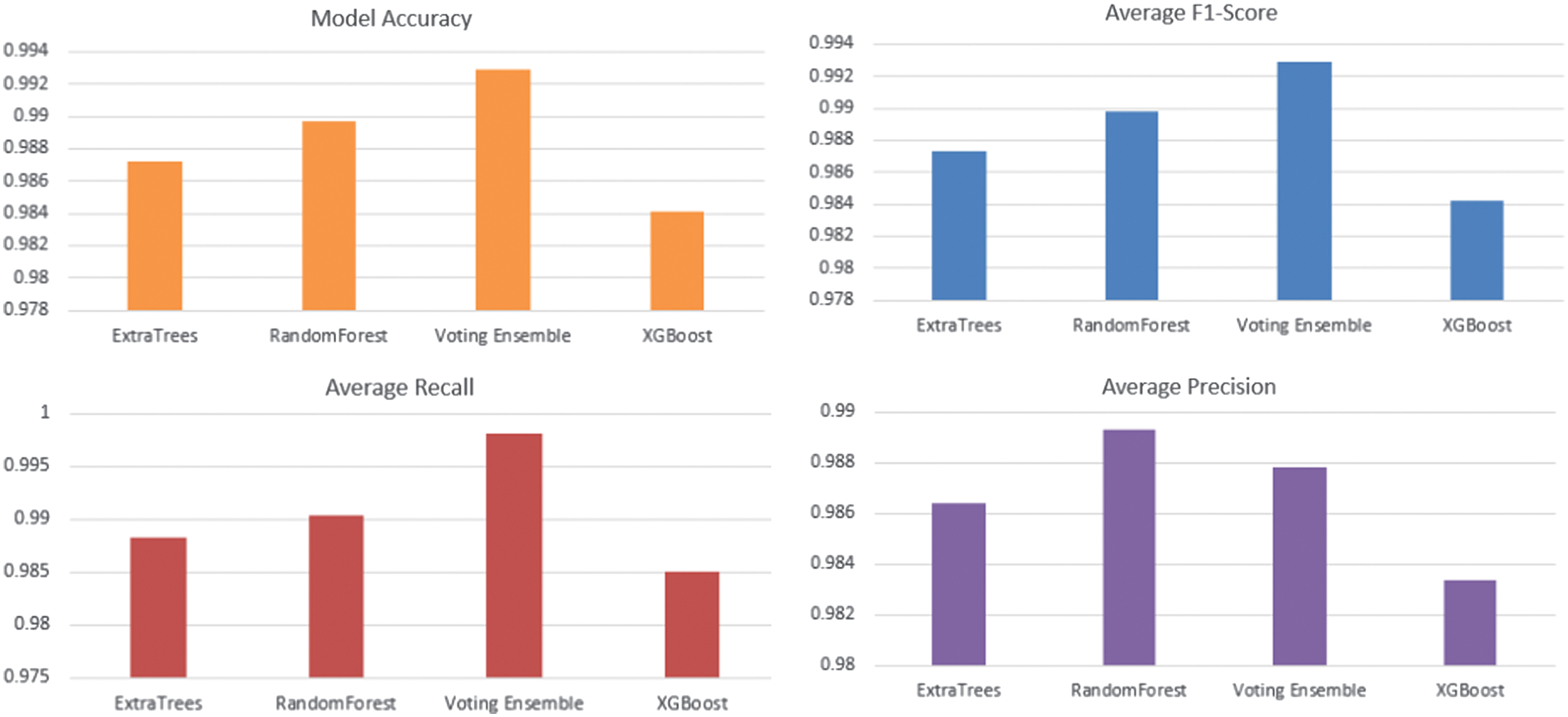 Fig. 2. Performance comparison of RandomForest, ExtraTrees, XGBoost, and the proposed Voting Ensemble in terms of classification metrics.
Fig. 2. Performance comparison of RandomForest, ExtraTrees, XGBoost, and the proposed Voting Ensemble in terms of classification metrics.
To demonstrate interpretability, we analyze the stability and frequency of the channels and regions selected by the proposed region-aware GA. The final selected feature set consists of eight EEG channels—FP1-F7, FP2-F8 (Frontal), C3-P3, C4-P4 (Central-Parietal), FT10-T8, FT9-FT10 (Temporal), and P7-O1, P8-O2 (Occipital)—along with four demographic features (Age, Gender, Medication Status, and Seizure History)
Figure 2 shows the performance analysis of the models of developed machine learning models of detecting epilepsy. Our presented Voting Ensemble model proves to perform much better and exhibits the greatest overall accuracy of 0.9929. When it comes to the most important task of seizure event detection (class 1), the ensemble returns the best F1-Score of 0.9928. This score implies that there is a real-life balance with respect to high precision (0.9980) and recall (0.9876), reducing false alarms and missed detections. Although some of the models such as RandomForest and XGBoost performed adequately, the combination of the models to form an ensemble classifier was much better and more reliable, and thus it formed the most appropriate model to be used to detect seizures automatically using EEG data. Figure 3 displays confusion matrices for four different machine learning models: Voting Ensemble, RandomForest, ExtraTrees, and XGBoost. Each matrix visualizes the performance of a classification model by showing the counts of true positive, true negative, false positive, and false negative predictions. For instance, in the ‘Voting Ensemble’ matrix, 9997 instances were correctly classified as class 0 (true negatives) and 9861 as class 1 (true positives). The off-diagonal values (19 and 123) represent misclassifications. Comparing these matrices with the previously discussed Precision, Recall, and F1-Score data, we can see how these metrics are derived from these counts, providing a comprehensive view of each model’s predictive accuracy and error types. The models generally show high accuracy, as indicated by the large numbers along the main diagonal.
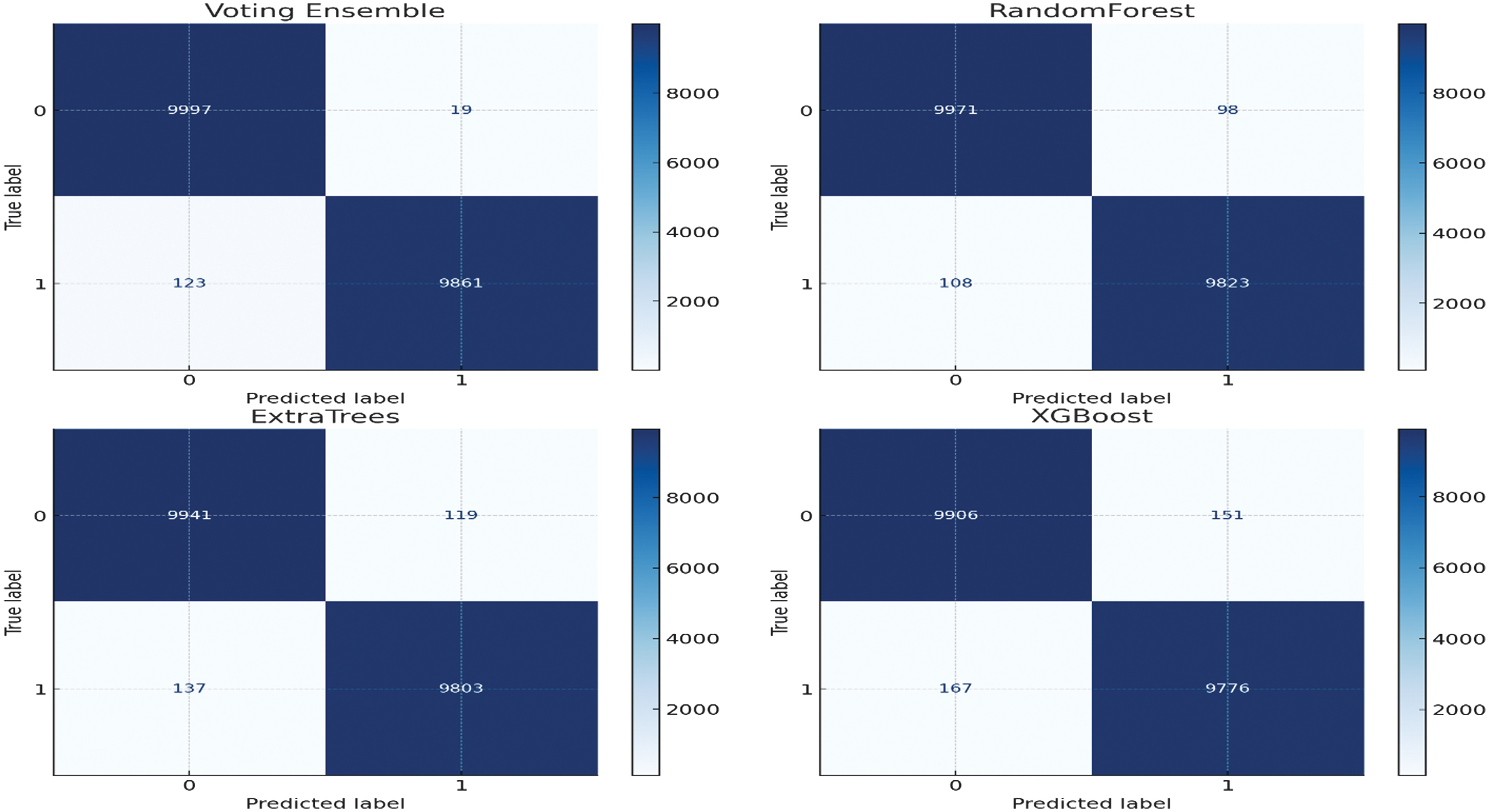 Fig. 3. Confusion matrices of all evaluated models showing class-wise prediction performance on the test set.
Fig. 3. Confusion matrices of all evaluated models showing class-wise prediction performance on the test set.
To assess robustness, all models were evaluated using a 70/10/20 train–validation–test split with 5-fold cross-validation, and the results are reported as mean ± standard deviation. The proposed Voting Ensemble achieved an accuracy of 99.29% ± 0.10%, demonstrating highly stable performance across folds. In contrast, RandomForest (98.97% ± 0.25%), ExtraTrees (98.72% ± 0.32%), and XGBoost (98.41% ± 0.37%) exhibited higher variance. The consistently lower standard deviation of the proposed method indicates superior robustness and reduced sensitivity to data partitioning. Statistical significance testing further confirms that the improvements of the proposed ensemble over the baseline models are not due to random variation (p < 0.05).
To quantify the contribution of the proposed region-aware GA feature selection, we performed an ablation study as shown in Table IV by replacing it with a conventional global GA that selects features from all EEG channels without considering neurophysiological regions. The global GA-based model achieved an accuracy of 89.5%, whereas the proposed region-aware GA-based framework achieved 99.29% ± 0.10%. This represents an absolute improvement of approximately 9.8%, demonstrating that incorporating brain-region structure into the feature selection process is crucial for both performance and robustness. These results confirm that the region-aware GA is a key component responsible for the superior performance of the proposed framework.
| Method | Feature selection strategy | Accuracy (%) |
|---|---|---|
| Global GA + Ensemble | Global (no region constraint) | 89.5 |
| Proposed method | Region-aware GA + Ensemble | 99.29 ± 0.10 |
1).COMPARISON
The various approaches to automated seizure detection are compared in Table V. The table outlines the feature processing techniques, classifiers, performance, and significant drawbacks associated with the most common methods in the literature. The specified framework, based on a new approach to feature selection, has multiple benefits in comparison to other existing methods. Its main advantage occurs in the feature selection process, which operates while being aware of the region that provides physiologically meaningful features that enable clinicians to learn which brain areas are contributing to a prediction and which are impossible to know using a non-interpretable end-to-end deep learning model. The model gets to be meta-aware as it will integrate patient demographic data, which will make it capable of wider generalization of various patient groups. Such specific feature suppression generates a small set of data that minimizes the computing costs involved. The system predicts behavior changes based on the selected features, which are very informative; therefore, the time series are processed and analyzed on a computer rather than on special equipment. As a result, our approach provides the best current accuracy in an efficient, implementable, and open way, thus making our approach a perfect solution to real-world clinical settings.
Table V. Comparison of existing seizure detection methods
| Approach | How it selects/processes features | Classifier | Accuracy | Drawbacks |
|---|---|---|---|---|
| XGBoost with hand-crafted EEG features (Random Cross-Validation (RCV) vs Leave-One-Out (LOO) comparison) [ | Extracts 53 handcrafted time and frequency domain features from 5s EEG windows after filtering | XGBoost with Optuna tuning | 82% (RCV) | Poor generalization to new patients; overfitting due to patient overlap in RCV |
| PANN with spatio-temporal EEG augmentation (STEA)[ | Learns patient-invariant features using adversarial training and synthetic seizure EEG generation | Patient-Adversarial Neural Network (PANN) | 95% | Requires complex adversarial training and synthetic data modeling |
| Hybrid CNN/DNN with PCA and SVM [ | Uses PCA for dimensionality reduction before SVM classification on CNN/DNN features | CNN-SVM-PCA, DNN-SVM-PCA | 99.42% | PCA may lose spatial info; the model may still be dataset-specific despite high accuracy |
| Hurst exponent + DWT-based EEG feature extraction with ML/DL models [ | Uses Hurst exponent, Daubechies-4 wavelet transform, ANOVA, and Random Forest (RF) regression for minimal feature selection on single-channel EEG | SVM, RF, Long Short-Term Memory (LSTM) | Up to 97% | Limited to single-channel data; handcrafted features may miss complex spatial patterns |
| Classical ML with 8 feature subsets and combined features [ | Evaluates multiple handcrafted EEG feature subsets and their combinations | Random Forest Classifier (RFC), SVM, K-Nearest Neighbors (KNN) | RFC (97.2%) | Limited to predefined features; performance varies by algorithm; Naive Bayes Classifier (NBC) underperforms (Area Under the Receiver Operating Characteristic Curve (AUROC) 0.729) |
| Hybrid XGBoost + RNN (LSTM/Gated Recurrent Unit (GRU)) for seizure prediction [ | EEG signal preprocessed; models trained on same dataset for binary classification | Hybrid XGBoost + LSTM/GRU | 98.20% | High computational cost; model complexity may hinder low-resource deployment |
| EfficientNet-B0 CNN + SVM ensemble with Short-Time Fourier Transform (STFT) and channel correlation [ | Extracts STFT and inter-channel correlation features from EEG; normalized preprocessing | EfficientNet-B0 + 6-SVM voting ensemble | 96.12% | Increased model complexity; ensemble may require careful tuning; potential latency for real-time use |
| RRWM and GMM features on CHB-MIT with KNN [ | Extracts RMS of RMS, mean absolute value, waveform length (RRWM), and GMM features from EEG | KNN (with hyperparameter tuning) | 99.03% | Limited to handcrafted features; tested on fixed window (patient-specific); lacks deep modeling |
| Mel-Frequency Cepstral Coefficients (MFCC) + Linear Predictive Cepstral Coefficients (LPCC) feature extraction with CNN-LSTM for seizure prediction [ | Converts EEG to Mel frequency domain, extracts MFCC and LPCC features to capture temporal-frequency changes | CNN-LSTM (best), compared with SVM, KNN, RF, Decision Tree (DT), Naive Bayes (NB), Linear Discriminant Analysis (LDA), Logistic Regression (LR) | 94% | Non-stationary signal variations still pose a challenge; Mel features may miss the spatial brain-region context |
VI.DISCUSSION AND LIMITATIONS
These findings provide firm support of our hypothesis that a region-aware and meta-informed strategy can mobilize state-of-the-art performance with regard to clinical interpretability. Our framework bridges the gap between computationally expensive black-box deep learning models and simpler, less context-aware traditional machine learning methods. Our method has several major strengths. It is very clinically interpretive since in contrast to global algorithms such as PCA that do not carry the physiological meaning of the feature in question, our region-wise GA will enable a clinician to visualize which part of their brain contributed to a prediction. The model further shows improved generalization through demographics of such patients, and hence by adding extremely valuable context, this model is effective in performing on a range of diverse patients. Moreover, the last model is very computationally efficient and runs on a small set of only 12 features, so the overhead is minimal compared to deep learning models and can be used in low consumption resources clinical environment. Lastly, our ensemble model also attains state-of-the-art accuracy of 99.28% to rival and even surpass numerous complex deep learning frameworks without compromising the transparency. This work shows that feature engineering that uses an intimate understanding of the domain can make a strong alternative to brute-force deep learning. How can we make this not only predictive but informative as well? The answer lies in building a model that, by insisting that the feature selection process respects the topology of the brain, is somehow informative. Despite the success of this study, it has a few limitations that can be exploited in further studies. The model has been built and tested on just a single, albeit extensive, dataset; its operation needs additional confirmation on other independent databases of various clinical organizations so as to determine its efficacy, as well as robustness. Also, the GA-based feature selection is done only once and a dynamic method, which might optimize the features selected depending on incoming patient information, may not be inferior to performance in the long run. Lastly, although we used four meta-features that were most predictive among clinical data, there exist other clinical data that could serve the model with better predictive capability, for example, the type of seizure or anti-epileptic drug, which could be added to the model.
VII.CONCLUSION
This paper has proposed a new framework for the detection of epileptic seizures, which was based on a special mix of both area-sensitive genetic feature selection, meta-integration of demographic data, and ensemble learning. The primary benefits of our approach are the avoidance of problems related to previously existing strategies, where aspects of clinical interpretability are either abandoned or their usefulness ignored. The suggested pipeline is smart so that it diminishes the dimensionality of EEG information 22 channels to 8 optimal, region-wise channels that are afterward supplemented with 4 patient-specific meta-features. The last soft-voting ensemble classification system demonstrated an excellent high-accurate rate of 99.28 on a very large reality-based dataset. This work shows that a domain-informed approach to building a machine learning pipeline can keep up with the performance of deep learning models in a black-box setting without their serious drawbacks and identify the key advantages of transparency, computational efficiency, and core clinical relevance. Our framework is an important milestone in the process of creating reliable and feasible AI-based applications helping to diagnose epilepsy and manage it. The novelty presented by this method is based on a peculiar synergy between the methodology, data processing, and design of the model. In essence, whereas global feature selection techniques such as PCA are blind to brain anatomy, our approach resorts to a region-wise GA, whereby not only the features will turn out to be optimal but also that they are meaningful at the neurophysiological level. This is a key difference from common deep learning models, which require raw EEG signals and substantial computational resources. Nonetheless, the model we propose performs well on tabular, processed data that is simple with a lightweight ensemble. Moreover, it enhances the study through the inclusion of patient demographic metadata which is not common in other researches. This trade-off enables our framework to overcome the fatal trade-off in the domain: it achieves state-of-the-art accuracy with the retention of high interpretability, which is critical toward clinical trust.
VIII.RECOMMENDATION
This research will leave behind a few major areas which are the subjects of future research. First, we will test the generalizability of the framework by conducting the validation on big multi-centric datasets. Second, we will discuss how to incorporate more sophisticated meta-features and how to explore dynamic feature selection. Last, we are working on compiling the model into a software tool that is user-friendly so that it can be used in real-time testing and deployment and will make this easy-to-implement solution and more accessible to neurologists.
IX.LIMITATIONS
We acknowledge that strict subject-wise evaluation using Leave-One-Subject-Out (LOSO) or group-based cross-validation is a stronger protocol for assessing cross-patient generalization; therefore, incorporating such subject-independent validation on multi-center datasets is identified as an important direction for future work.