I.INTRODUCTION
Traumatic brain injury (TBI) is a critical population health problem that is estimated to impact millions of individuals per year and is one of the most common causes of death and disability in the global population. Globally, a total of more than 69 million people experience TBI annually, and the distribution of the issue is incredibly skewed. TBIs in developed countries can be linked to sport injury, falls among the elderly, and road traffic accidents, whereas road traffic accidents continue to be the leading cause of TBI prevalence in low- and middle-income countries. It is important to note that TBI has enormous financial costs in the short term (direct clinical consequences) and the long term (indirect consequences) affecting the family, healthcare delivery systems, and the economy through cognitive, physical, and psychosocial effects. Chronic impairments in attention, memory, and motor coordination are common in survivors of TBI, which decreases their quality of life and leads to chronic dependence. These dimensions make the accurate diagnosis, timely treatment, and successful classification of TBI subtypes essential not only to the fate of individual sufferers but also to the reduction of the general impact of the disorder on society.
Intracranial Hemorrhage (ICH) is one of the most clinically significant consequences of TBI and occurs when blood gathers in or around the brain due to traumatic injuries to blood vessels. ICH does not exist as an entity, but it is a continuum of various categories of hemorrhages, some of which include epidural, subdural, subarachnoid, intraparenchymal, and intraventricular hemorrhages. Each of these subtypes implies a different clinical implication. When it comes to skull fractures, the epidural hemorrhages can grow and require urgent neurosurgical intervention. Subdural hemorrhages, in turn, may proceed at a slower pace, yet they are also associated with a high mortality rate in the case of failure to treat them. Subarachnoid bleeding is associated with the risk of vasospasm and delayed ischemia, and intraventricular bleeding could lead to the blockage of cerebrospinal fluid channels and consequently to hydrocephalus. The intraparenchymal hemorrhages that occur in brain tissues may cause devastating focal deficits depending on their localization. In light of such multiplicity, the appropriate categorization of ICH subtypes is critical to selecting the appropriate clinical pathway and prioritizing interventions in acute care.
Non-contrast computed tomography (CT) is the standard imaging modality in modern clinical practice, since it is employed to detect the existence of intracranial hemorrhage within the shortest time possible. CT is associated with a number of advantages: it is standard, it is less expensive when compared to Magnetic Resonance Imaging (MRI), and the images they depict can be delivered very fast when faced with an emergency in an acute setting. Acute blood is very sensitive to CT and, thus, it is very useful in trauma treatment where time is a very important consideration. Reliance on CT is problematic in itself, however. First, radiologist interpretation is never a quick process and can be prone to inconsistency as it can be influenced by human factors. In busy emergency departments where the radiologist must attend to huge caseloads, there is an increased risk of delayed diagnosis. Second, inter-observer variability exists, especially with the mild or marginal cases. In other words, a small subarachnoid hemorrhage may be missed during one examination and identified during another. Finally, the availability of qualified and prompt evaluation is restricted due to a large gap in the number of trained neuroradiologists, particularly in low-resource settings. These weaknesses indicate that there is an acute necessity to create augmented diagnostic devices that can help radiologists improve the speed and accuracy of hemorrhage classification.
In recent years, machine learning (ML) and artificial intelligence (AI) have become the disruptive technologies in the medical imaging industry, which can answer the majority of these issues. AI models have already been shown to be useful in other domains of TBI management, such as image processing via computer and outcome prediction. They can be explainable ML models such as E-TBI that can provide interpretable patient post-TBI outcome predictions that guarantee the decision-making process is both transparent and clinically meaningful [1]. A different study also proved that ML can be effective in forecasting when emergency neurosurgery is required within 24 hours of severe or moderate TBI and offer clinicians real-time decision support tools in life-threatening scenarios [2]. Biomarker-based diagnostics have also been used in conjunction with computational models where proteins were quantified to determine the extent and course of injury, such as glial fibrillary acidic protein (GFAP) and ubiquitin carboxy-terminal hydrolase L1 (UCH-L1) [3]. These examples represent the scope of AI implementation in TBI and how it could be applied to integrate imaging, clinical, and molecular data and deliver holistic patient care.
The use of deep learning models has brought a revolution to the automated interpretation of CT scans in the imaging field. They have been shown in validation studies to achieve radiologist level in bleeding and injury severity grading [4]. Besides detection, deep learning models have also been applied to other secondary injury mechanisms, such as the role of mitochondrial dysfunction in propagating inflammation and cell damage following TBI [5]. These understandings would extend beyond the short-term diagnosis to help clinicians understand the biological foundations of long-term sequelae. It is worth noting that AI-based solutions have been demonstrated to be applicable in other healthcare settings. It is established that predictive frameworks can be highly generalizable in low-resource and high-income environments, implying that they may help to decrease inequality in access to health worldwide [6]. The additional new trends include non-invasive diagnostic assays, for example, acoustofluidic separation as a means of identifying TBI biomarkers in circulating exosomes [7], and the use of ML models to conduct multi-class imaging classification [8].
Going a step further, researchers have examined the application of AI to pediatric groups, wherein TBI poses distinct diagnostic and management issues. Indicatively, neural network-related solutions have been suggested to identify mild TBI in children, emphasizing the adaptability of AI models to the cohort specializations [9]. Moreover, ML-based mortality prediction models have offered patient prognosis assessment tools and have been applied to intensive care and trauma patients [10]. Collectively, these papers indicate that AI is not limited to one area of TBI research but cuts across imaging, biomarkers, patient outcomes, and prognostic modeling.
In spite of these developments, however, there is still a massive gap in research: no strong frameworks of fine-grained, multi-classification of subtypes of intracranial hemorrhage are available. Most of the published literature has concerned binary classification, for example, hemorrhage versus non-hemorrhage, or outcome prediction, for example, mortality or surgical requirement. Although such methods are clinically helpful, they fail to assist in the more subtle problem of discriminating between subtypes of hemorrhage, each of which has its own prognostic and therapeutic consequences. As an example, it is necessary to distinguish between subarachnoid and intraventricular hemorrhages, as they require different management approaches, but this distinction is poorly studied with automated systems. This is a limitation that explains why there is an urgent need to develop models that transcend binary classification and address the multi-class labeling challenges in CT scans.
To fill this knowledge gap, the current study proposes the Swin Transformer, which is a new Vision Transformer (ViT) architecture that can be optimally trained to achieve a trade-off between computational efficiency and high performance in image classification. Conventional convolutional neural networks (CNNs) have been the dominant medical imaging technology since they can capture local patterns. CNNs are, however, naturally constrained by their fixed receptive fields that restrict their capacity to capture long-range interdependencies and global context. By comparison, ViTs perceive images as a sequence of patches and use self-attention to learn relationships on a whole image. This makes it possible to extract rich features and perform better in context-dependent activities. The Swin Transformer also builds upon the unstructured ViTs by implementing hierarchical windowed self-attention; that is, the calculation of attention is carried out in local windows to minimize computational cost. To maintain that global context, the architecture has used shifted windows which permit information to spread in various parts of the image. This is the best of both worlds: efficient computation and global receptive fields.
The novelty of this study is in its use of the Swin Transformer for the problem of multi-classification of subtypes of intracranial hemorrhage, thereby further extending the binary detection paradigms, which currently dominate the literature. In the following way, we can summarize our contributions. We create a balanced dataset of 60 000 CT images using a 1-to-1 ratio of six classes: epidural, intraparenchymal, intraventricular, subarachnoid, subdural, and normal. This ensures that the model is trained in an equal representation of classes, and this will minimize the bias that natural imbalances in real-world data introduce to the model. Second, we conduct Swin Transformer training on this dataset using a highly developed training program that includes optimization schemes such as AdamW, cosine annealing scheduling, and label smoothing to reach the highest generalization. Third, we conduct stringent statistical validation, including K-fold cross-validation to test stability and the McNemar test to test statistical significance compared to a strong base model (EfficientNetV2-S). Finally, explainability techniques are applied as well, though through Grad-CAM++, to ensure that one of our models bases their decision-making on clinically relevant image regions.
In thus doing, this work establishes a new standard in the multi-class ICH classification, not only in quality but also in the rigor of validation. They demonstrate both state-of-the-art performance and statistically robust, interpretable performance, which are essential in clinical translation. The study is itself a contribution to a gap in the existing literature, since it does not stop at binary detection tasks and fine-grained classification, which are the foundations of future AI-based neurotrauma care devices.
The rest of this paper is organized in the following way. Section II conducts a literature review of pertinent literature in the area of biomarkers, imaging, and ML models and explainable AI. In Section III, the dataset, preprocessing pipeline, Swin Transformer architecture, and the experimental setup are provided. The results are reported in Section IV (quantitative measures, statistical validation, and explanation tools). A discussion of the main contributions, clinical implications, and future research directions is provided in Section V.
II.LITERATURE REVIEW
Using AI in TBI research has evolved on many fronts, since it is a complicated disorder and its clinical presentations can have different manifestations. The first neuroimaging research is on functional and metabolic imaging, and researchers started investigating how advanced imaging technologies can be used to detect the functional neural perturbations. Vedaei et al. [11] reported that resting-state functional MRI (rs-fMRI) and ML methodology could improve the discrimination of chronic mild TBI and therefore indicated the potential of multimodal imaging features to unveil the patterns of injury that are likely to pass through the usual CT scans. These papers have shown the relevance of deep learning to detect the biomarkers of connectivity disruption, which is subtle and challenging to identify through human experts. The other significant limitation in these works, however, is that the more complicated a model is, the less the decision-making process could be understood, which brought up the question of interpretability and clinical acceptability.
This is what catalyzed explainable AI in TBI research. To address this, Ngo et al. [12] suggested the E-TBI system, which combined predictive modeling with interpretability features that allow a clinician to understand and trust the algorithmic decisions made in the system. Consistent with that, the explainable ML employed by Liu et al. [13] in predicting sepsis in TBI patients proves transparency to be an important factor in high-stakes clinical predictive processes. Explainability has become a broader trend in medical AI: performance metrics such as accuracy or recall are not sufficient without interpretability mechanisms that are clinically consistent. Such congruency is especially topical in the context of TBI, where the treatment decisions might be life or death.
In addition to explainability, biological and cellular markers have emerged as the new target interest of the TBI research, which offers a possibility to extend the imaging-only diagnostics. Surrogately, extracellular mitochondria have been shown to mediate post-TBI-induced acute lung injury to interrelate neural trauma and systemic inflammatory pathways [5]. It has been proposed as well that NINJ1-mediated endothelial rupture is a predisposing factor to the escalation of blood–brain barrier destruction after TBI, which also plays a role in accelerating the development of the neurological failure [12]. These findings restate that TBI is systemic and that secondary injuries in peripheral organs and microvascular damage within the brain exacerbate this primary trauma. Subsequent pediatric-based reviews followed this study by Lampros and co-authors [13] who concluded that the biomarkers (GFAP and UCH-L1) had special potential in the early diagnosis of susceptible groups. Additional studies by Kim et al. [14] estimated that AI-enhanced neurocritical care systems would use these biomarkers as part of continuous monitoring pipelines, which feed clinicians with real-time feedback. In the same vein, Yao et al. [15] have performed a systematic review of the prognostic applications of ML in mild TBI and have shown how biomarker-based models will guide individual rehabilitation decisions.
Simultaneously, advances in biosensing technologies have driven progress toward point-of-care diagnostics. Özcan et al. [18] developed a quartz tuning fork (QTF) biosensor for GFAP detection, providing an innovative, mass-sensitive approach for rapid screening. Tan et al. [32] introduced surface-enhanced Raman scattering assays for detecting UCH-L1, while Gao et al. [34] designed an ultrasensitive lateral flow assay for S-100β, a well-established marker of TBI. Legramante et al. [33] further demonstrated that GFAP and UCH-L1 assays could triage mild TBI cases by identifying patients unlikely to require CT scans, thereby reducing unnecessary radiation exposure. Collectively, these studies highlight a translational shift toward bedside diagnostics, though their direct application to hemorrhage subtype classification remains indirect, focusing instead on severity stratification and triage.
Although the biomarker approach has emerged, CT imaging remains the clinical gold standard in the evaluation of acute TBI, especially in the case of intracranial hemorrhage. Several experiments have shown that deep learning on the raw CT data is both possible and more precise. Jiang et al. [4] confirmed the use of a deep learning model to detect TBI and Neuroimaging Radiological Injury Score (NIRIS) grading in a multi-reader environment with promising results and challenges in identifying small lesions. By combining CT imaging with bioclinical markers at presentation, Mekkodathil et. al. [16] were able to predict mortality better, and it is proposed that the combination of multimodal data streams will help improve outcome estimates. Uparela-Reyes et al. [17] demonstrated the expanded use of AI in TBI imaging through a bibliometric analysis that demonstrated an exponential increase in the number of publications over the last 10 years, as the world began to become interested in the use of computational models. They also pointed out, however, that to date, the majority of AI models have not moved beyond research to routine clinical practice, and that validation, generalizability, and acceptability to clinicians are all problematic.
Another promising area has been segmentation-based methods, which allow to accurately measure the volume of hemorrhage. A CT-based hemorrhage segmentation system is reported by MacIntosh et al. [21] and has been demonstrated to be useful in both stroke and TBI. However, their results also showed limitations in extrapolability to other scanners and imaging protocols, which is a common problem with AI when it comes to radiology. Synthetic reviews by Lin and Yuh [22] and Gudigar et al. [23] summarized the set of computational strategies used to process TBI imaging, such as CNNs, feature-based, and hybrid. The two reviews stressed that AI models are always associated with a high level of performance on internal datasets, and external validation is poorly represented. Wu et al. [24] further applied this trend to outcome prediction in severe TBI and demonstrated that ML performed better as compared to conventional statistical methods to estimate mortality. These initiatives demonstrate the opportunities of AI to supplement radiological knowledge but also warn against overinterpretation because it has not been multi-institutionally validated.
Prognostic modeling has also become a concomitant research area, aiming to forecast secondary outcomes (hematoma progression, systemic complications, and survival). Mohammadzadeh et al. [25] performed a systematic review and meta-analysis demonstrating that ML could be used in relation to predicting hematoma progression, which is one of the most crucial factors to determine its prognosis. Phaphuangwittayakul et al. [26] took this step further and elaborated a deep learning method of multi-lesion quantification, associating the identification of lesions with the quantification of their volumes. The clinical usefulness of computational methods is proven by a meta-analysis by Courville et al. [27], who found ML algorithms to be superior to traditional models at outcome prediction. The scope of this is extended by Peng et al. [28], who forecasted severe acute kidney injury in patients with TBI, and Ge et al. [31], who simulated sepsis-related acute brain injury, both of which explored how systemic complications might be predicted through AI models. Critical comments on biomarker-based prognostication and the problems with reproducibility in neuroimaging studies were made in complementary reviews by Ghaith et al. [29] and Pierre et al. [30], respectively.
Biosensing is an ongoing parallel innovation that continues to expand the possibilities of prognostics. Tan et al. [32] have shown that Raman scattering-based assays can also detect UCH-L1 with high sensitivity, providing the potential to conduct early prognostic triage. In an article by Legramante et al. [33], the authors contrasted the results of their tests, which included GFAP and UCH-L1, with CT exclusion protocols, and emphasized their utility in eliminating unnecessary imaging in patients with mild TBI. Gao et al. [34] reported an enhanced lateral flow assay of S-100 2 that expands the limits of portable, ultrasensitive biomarker detection. Nevertheless, despite these technological advances, Hibi et al. [35] raised some concerns about the preparedness of existing automated CT-based models to be integrated into clinical practice by citing limitations in dataset diversity, algorithm stability, and understandability.
In these diverse directions, a somewhat homogeneous strand has been located: the increasing prominence of hybrid and explainable architectures. Singh and Rakhra [8] have also emphasized the integration of imaging and outcomes into single AI pipes and how this could mediate between diagnostic and prognostic processes. Lampros et al. [13] emphasized that a child population has specific needs since TBI is linked to various diagnostic and treatment problems in this patient group. Kim et al. [14] approximated that in the future, AI-enhanced neurocritical care would be introduced into the world, which will encompass continuous imaging, biomarkers, and clinic data to develop full-scale decision-support systems. Of paramount interest has been the explainability of such innovations. In the works of Ngo et al. [12], Liu et al. [13], and others, the key to clinician trust is transparent models. Gradient-weighted Class Activation Mapping (Grad-CAM) and SHapley Additive exPlanations (SHAP) are only two examples of methods that have been applied to characterize the decisions made by neural networks, so as to align the algorithmic truth with radiological judgment, and to address the black-box prediction/clinical explanation gap [27].
Combined, the literature reviewed indicates significant progress in AI in TBI in the areas of imaging, biomarkers, prognostication, and biosensing. Nevertheless, it also indicates an underlying and long-standing research gap: the multi-classification of subtypes of hemorrhage in CT scans has not been well researched. Current available work is strongly biased in detecting binaries or prognosticating outcomes, and the immediate diagnostic task of distinguishing epidural, subdural, subarachnoid, intraparenchymal, intraventricular, and normal scans is significantly underexploited. Although biomarkers and prognostic models guide secondary management, the basis on which treatment strategies are developed is the acute classification of the subtypes of hemorrhage. In this work, the Swin Transformer is implemented to fill in this gap by deploying a hierarchical ViT architecture to realize effective multi-class hemorrhage classification with stringent statistical validation and explainability. By placing our work in this more general flow of AI research, we are helping to move our field beyond binary and prognostic paradigms, to clinically actionable, fine-grained diagnostic support. [11–35].
Although hybrid CNN–Transformer architectures have shown potential in specialized clinical studies, most publicly available multi-class hemorrhage datasets report comparatively lower performance, typically in the 70–80% range. Achieving an accuracy of 82.02% in this work therefore represents an improvement over several prior transformer- and CNN-based approaches trained under similar data constraints. Higher accuracies reported in hybrid frameworks generally rely on private, richly annotated datasets and complex multi-branch models that limit reproducibility. As the availability of expert annotations improves, future extensions of this work will explore hybrid fusion models to further enhance classification granularity and performance.
Despite the significant progress achieved by CNN-based and hybrid deep learning models in intracranial hemorrhage detection, existing works remain limited in their ability to capture long-range spatial dependencies and global contextual relationships within CT images—factors that are critical for reliable subtype-level classification. Most prior studies also rely on imbalanced datasets, lack statistical validation, or provide limited interpretability, restricting their clinical applicability. Furthermore, only a few recent approaches have explored transformer-based architectures for medical imaging, and these are primarily focused on binary detection tasks rather than comprehensive multi-class categorization. This gap highlights the need for a unified framework that integrates global attention mechanisms, robust validation strategies, and explainable visualization to enhance diagnostic confidence. Building upon these observations, the next section presents the proposed Swin Transformer-based pipeline, detailing the dataset construction, architectural components, training strategies, and evaluation protocols employed in this study.
III.METHODOLOGY AND EXPERIMENT
A.DATASET DESCRIPTION
The dataset used in this study was self-collected from Shriman Hospital, Jalandhar, under the supervision of certified radiologists and following institutional ethical guidelines. The acquired CT scans contained six categories of intracranial conditions: epidural hemorrhage, intraparenchymal hemorrhage, intraventricular hemorrhage, subarachnoid hemorrhage, subdural hemorrhage, and normal scans. To address class imbalance and increase the dataset size, we applied structured augmentation techniques, including rotation, intensity variation, cropping, and Gaussian noise injection. These operations resulted in a balanced dataset of 60,000 CT slices, with approximately equal representation across all six classes. To support reproducibility and open research, the final augmented dataset has been publicly released on the Mendeley Data repository. No external hospital data or multiple-source aggregation was used in this study; all raw data originated from Shriman Hospital, Jalandhar. The dataset employed in this study comprised 60,000 head CT images, evenly distributed across six clinically relevant categories of intracranial hemorrhage: epidural, intraparenchymal, intraventricular, subarachnoid, subdural, and normal (ne). Each class contained 10,000 images, ensuring perfect balance and preventing class-specific bias. Representative CT slices for each class are shown in Fig. 1.
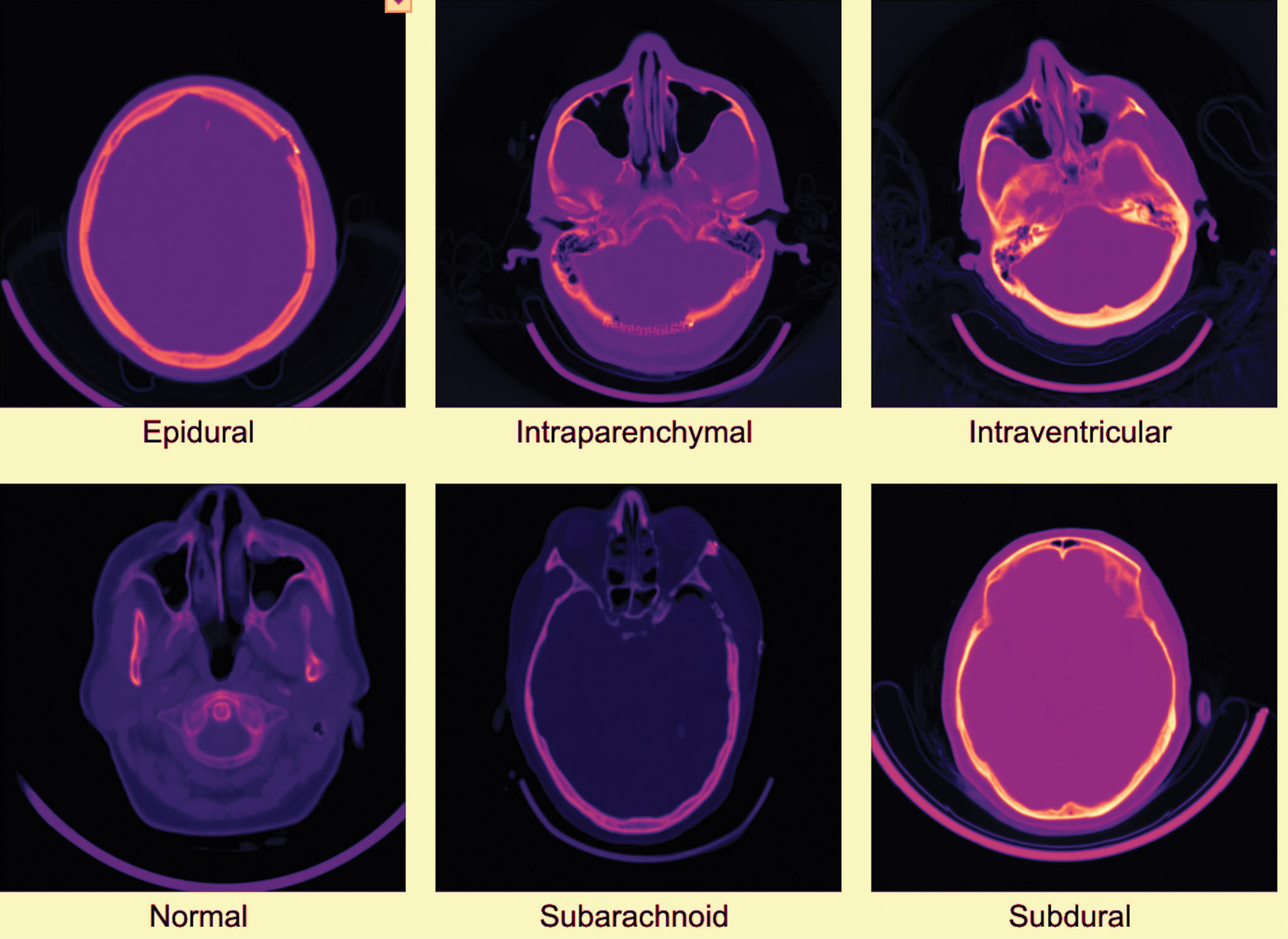 Fig. 1. Examples of head CT scans for each of the six classes in the dataset.
Fig. 1. Examples of head CT scans for each of the six classes in the dataset.
The dataset is partitioned using stratified sampling into 70% training (42,000 images), 15% validation (9,000 images), and 15% testing (9,000 images), with each subset maintaining the same 1:6 distribution across classes.
Although the dataset provides a balanced representation of six intracranial hemorrhage categories through structured augmentation, it is important to acknowledge certain limitations. The raw CT scans were collected from a single clinical center (Shriman Hospital, Jalandhar), which may limit the diversity of imaging characteristics such as scanner type, acquisition protocols, and demographic variability. Additionally, the dataset was not combined with external hospital repositories, which restricts the model’s exposure to broader real-world variations. Future work will incorporate multi-center data collection, larger radiologist-validated annotations, and more heterogeneous imaging sources to improve generalizability and clinical robustness.
B.DATA PREPROCESSING
All CT images were resized to 224 × 224 pixels to match the Swin Transformer input. Pixel values were normalized to [0,1]. Standard augmentations (random horizontal flips, random rotations, and contrast adjustments) were applied to improve generalization while preserving clinical fidelity.
C.MODEL ARCHITECTURE: SWIN TRANSFORMER
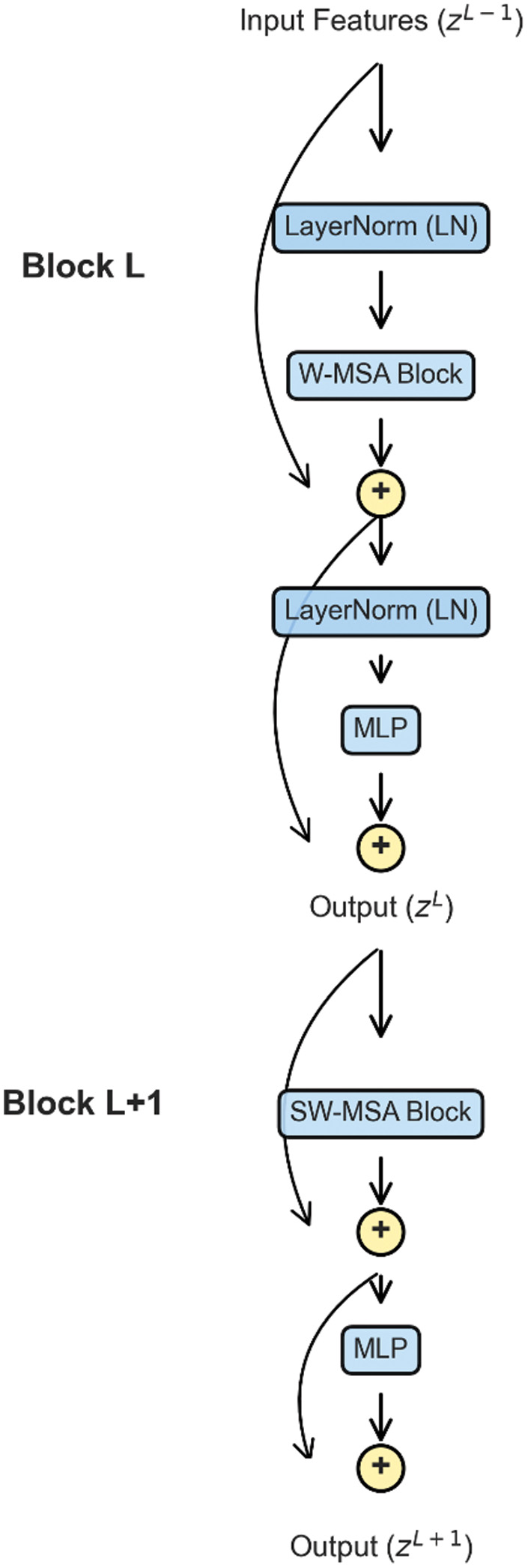
The backbone of this study is the Swin Transformer (swin_tiny_patch4_window7_224), a hierarchical ViT. Unlike conventional convolutional models, it leverages a windowed multi-head self-attention (W-MSA) mechanism to restrict computation to local windows, thereby reducing complexity, while the shifted window attention (SW-MSA) enables cross-window interactions to capture long-range dependencies and global context. The complete workflow of the proposed methodology—from dataset preprocessing through model training to final classification output—is illustrated in Fig. 2.
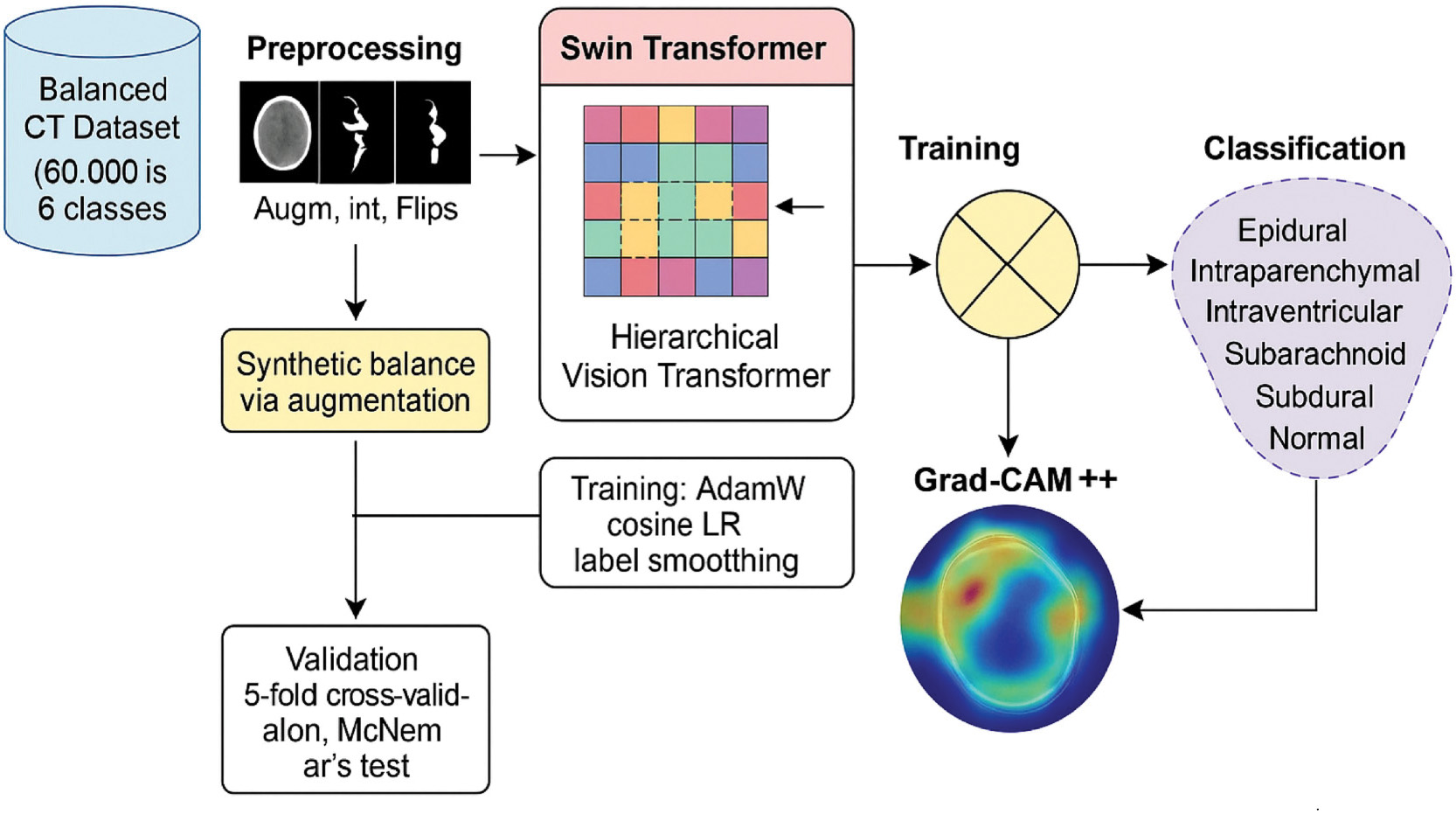 Fig. 2. Workflow of the proposed methodology from dataset preprocessing to classification output.
Fig. 2. Workflow of the proposed methodology from dataset preprocessing to classification output.
Mathematical Formulation
The scaled dot-product attention for a single head is computed as:
where Q, K, and V are the query, key, and value matrices derived from token embeddings, respectively, is the key dimension, and B is the relative positional bias.For an input sequence , the operations within consecutive Swin Transformer blocks are as follows:
Here, LN represents layer normalization, while MLP denotes a feed-forward neural network (Fig. 3).
D.TRAINING PROCEDURE AND HYPERPARAMETERS
The model is trained using PyTorch with the AdamW optimizer, a weight decay of 0.05, and an initial learning rate of 1e-4 scheduled by cosine annealing. Training is conducted for 40 epochs, with a batch size of 32. The loss function is cross-entropy with label smoothing (ɛ = 0.1) to prevent overconfidence in class predictions. The complete hyperparameter configuration is summarized in Table I.
Table I. Hyperparameter configuration for Swin Transformer
| Parameter | Value |
|---|---|
| Model | swin_tiny_patch4_window7_224 |
| Optimizer | AdamW |
| Learning rate | 1e–4 |
| Weight decay | 0.05 |
| Scheduler | Cosine annealing |
| Loss function | Cross-entropy (label smooth = 0.1) |
| Epochs | 40 |
| Batch size | 32 |
| Image size | 224 × 224 pixels |
E.ALGORITHMIC REPRESENTATION
Algorithm 1. Training Procedure for Swin Transformer
| 1. Load dataset and apply preprocessing (resize to 224 × 224, normalize). |
| 2. Initialize Swin Transformer (swin_tiny_patch4_window7_224). |
| 3. Define optimizer (AdamW), loss function (Cross-Entropy + Label Smoothing). |
| 4. For epoch = 1 to 40 do: |
| a. For each batch in training set: |
| i. Perform forward pass. |
| ii. Compute loss. |
| iii. Backpropagate gradients. |
| iv. Update parameters. |
| b. Evaluate model on validation set. |
| c. Adjust learning rate via cosine annealing. |
| 5. Save best model checkpoint (based on validation accuracy). |
| 1. Split D into K stratified folds. |
| 2. For each fold i: |
| Train Swin Transformer on K-1 folds. |
| Validate on fold i. |
| Record accuracy A_i. |
| 3. Compute mean μ = (Σ A_i)/K. |
| 4. Compute variance σ2 = (Σ (A_i - μ)2)/K. |
| 1. Initialize counts: b = 0, c = 0 |
| 2. For each sample: |
| if A correct and B incorrect → b++ |
| if A incorrect and B correct → c++ |
| 3. Compute statistic: |
| χ2 = (|b − c| − 1)2/(b + c) |
| 4. Compute p-value from χ2 distribution (df = 1). |
| 5. If p < 0.05 → reject null hypothesis. |
IV.RESULT AND DISCUSSION
A.QUANTITATIVE PERFORMANCE ANALYSIS
The Swin Transformer (swin_tiny_patch4_window7_224) achieved a test accuracy of 82.02%, demonstrating its effectiveness in classifying intracranial hemorrhage subtypes. A detailed classification report is presented in Table II, showing class-wise precision, recall, and F1-score.
Table II. Detailed classification report for Swin Transformer
| Class | Precision | Recall | F1-score | Support |
|---|---|---|---|---|
| Epidural | 0.9607 | 0.9453 | 0.9530 | 1500 |
| Intraparenchymal | 0.7775 | 0.7733 | 0.7754 | 1500 |
| Intraventricular | 0.8352 | 0.8580 | 0.8464 | 1500 |
| Normal | 0.8086 | 0.8027 | 0.8056 | 1500 |
| Subarachnoid | 0.7369 | 0.7600 | 0.7483 | 1500 |
| Subdural | 0.8062 | 0.7820 | 0.7939 | 1500 |
| Macro avg. | 0.8208 | 0.8202 | 0.8204 | 9000 |
| Weighted avg. | 0.8208 | 0.8202 | 0.8204 | 9000 |
The results indicate that the model performs exceptionally well in detecting epidural hemorrhage (F1 = 0.9530), while slightly lower performance is observed for subarachnoid hemorrhage (F1 = 0.7483), which is consistent with its more diffuse radiological features. The overall distribution of predictions across all six classes is visualized in the confusion matrix shown in Fig. 4, which highlights both the strengths and the remaining challenges in subtype classification.
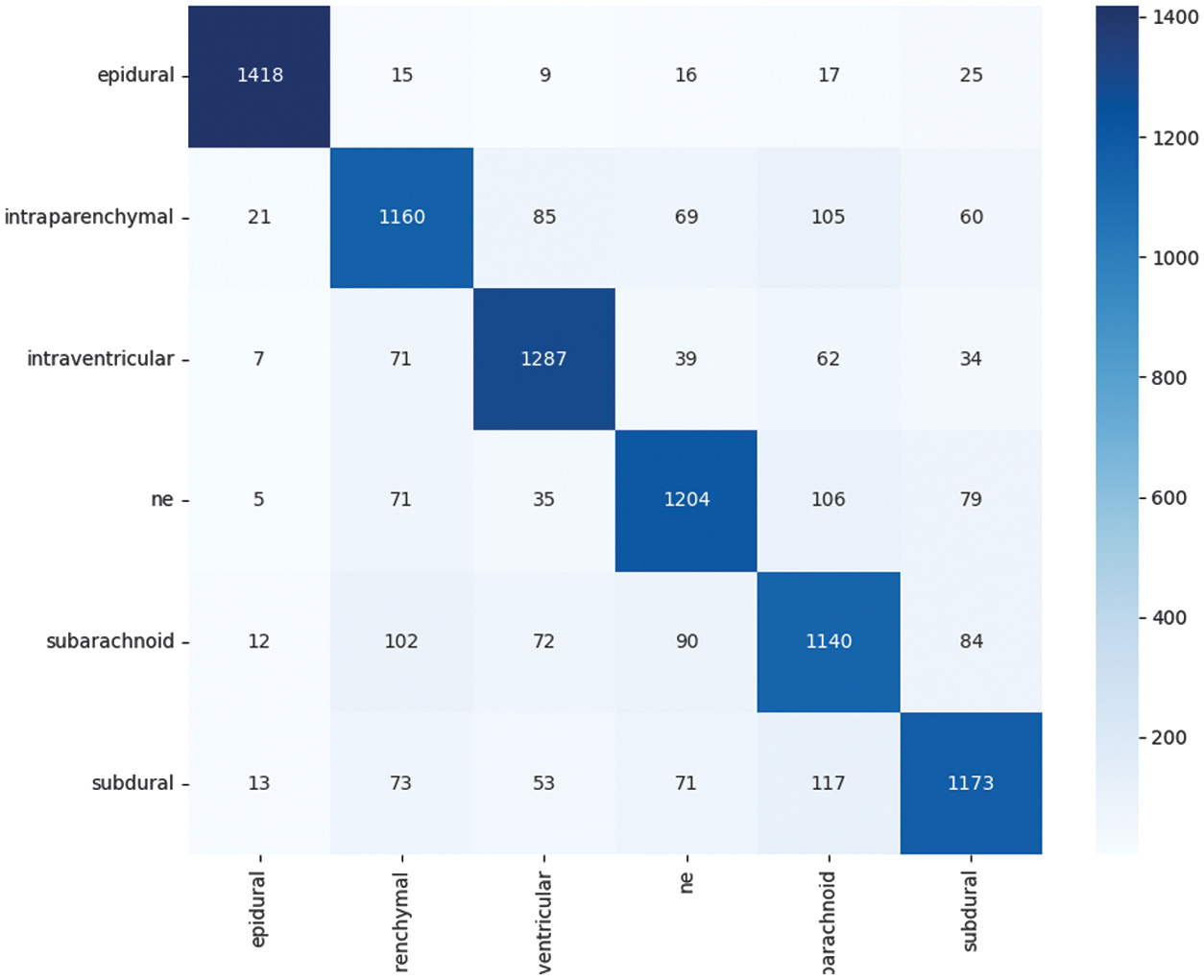 Fig. 4. Confusion matrix of Swin Transformer predictions on the test dataset.
Fig. 4. Confusion matrix of Swin Transformer predictions on the test dataset.
B.MODEL STABILITY (5-FOLD CROSS-VALIDATION)
To evaluate robustness, a 5-fold cross-validation is conducted on a 30% stratified subsample of the dataset, with each fold trained for eight epochs.
- •Individual fold accuracies: [57.47%, 58.44%, 57.31%, 55.44%, 57.64%]
- •Mean accuracy: 57.26%
- •Standard deviation: 0.99%
Although the mean accuracy is lower due to the reduced number of training epochs compared to the main experiment (8 vs. 40), the extremely low standard deviation demonstrates strong stability and consistency of the model across different subsets.
The stability is formally expressed as:
where represents the accuracy for the fold, is the mean accuracy, and is the standard deviation.For our case, and
This numerical demonstration proves that despite the shorter training cycle, the Swin Transformer exhibits high robustness and reliable performance across varying data partitions. The fold-wise accuracy distribution is summarized in Fig. 5, which visualizes the consistency of performance across all five cross-validation experiments.
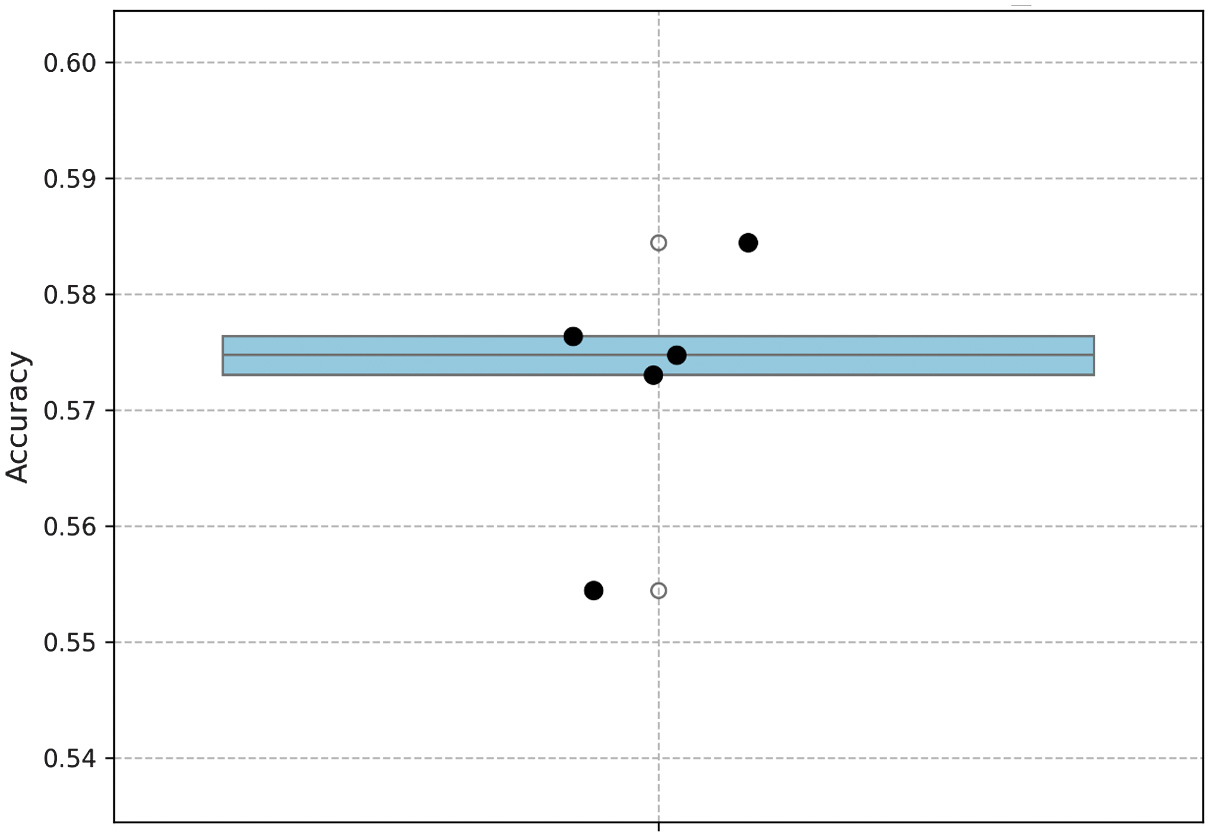 Fig. 5. Accuracy distribution across 5-fold cross-validation experiments.
Fig. 5. Accuracy distribution across 5-fold cross-validation experiments.
C.MODEL COMPARISON (McNEMAR’S TEST)
The performance of swin_tiny is statistically compared against EfficientNetV2-S, which had previously achieved an accuracy of 80.17%. To evaluate whether the observed performance difference is statistically significant, McNemar’s test is applied to the contingency of disagreements between the two models. The distribution of agreements and disagreements is presented in Table III, which forms the basis of the chi-squared calculation.
Table III. McNemar’s test contingency table of disagreements
| EfficientNetV2-S correct | EfficientNetV2-S incorrect | |
|---|---|---|
| Swin-Tiny correct | Agreement | 727 |
| Swin-Tiny incorrect | 560 | Agreement |
The test statistic is given by:
The corresponding p-value is given by:
Since the p-value is far below the threshold of , the null hypothesis is rejected. This confirms that Swin-Tiny is statistically superior to EfficientNetV2-S for intracranial hemorrhage classification, as illustrated in Fig. 6, which visualizes the outcomes of McNemar’s test comparison between the two models.
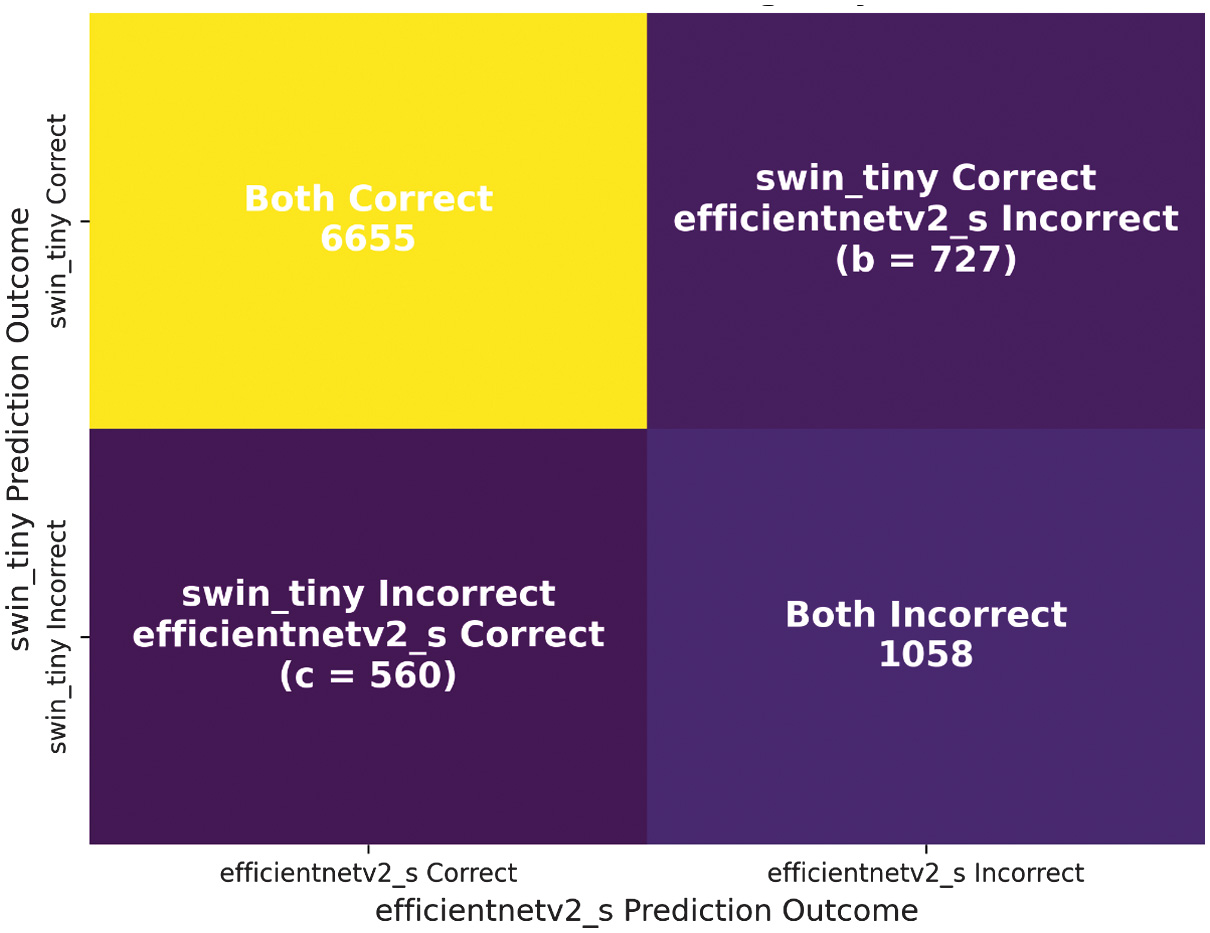 Fig. 6. McNemar’s test comparison between Swin-Tiny and EfficientNetV2-S.
Fig. 6. McNemar’s test comparison between Swin-Tiny and EfficientNetV2-S.
D.QUALITATIVE ANALYSIS (EXPLAINABILITY WITH GRAD-CAM++)
To interpret the model’s decision-making, Grad-CAM++ visualizations were generated for representative samples across all six classes. For clarity, each example is presented as a side-by-side comparison of the original CT scan and its corresponding Grad-CAM++ heatmap overlay. The visualizations confirmed that the model’s focus areas corresponded to clinically relevant hemorrhagic regions. For instance, in epidural hemorrhages, the attention maps highlighted sharply localized high-density regions near the skull, while in subarachnoid hemorrhages, diffuse cortical sulcal involvement is emphasized. This alignment between radiological features and model focus enhances the clinical interpretability of the framework. Representative examples of these comparisons are shown in Fig. 7.
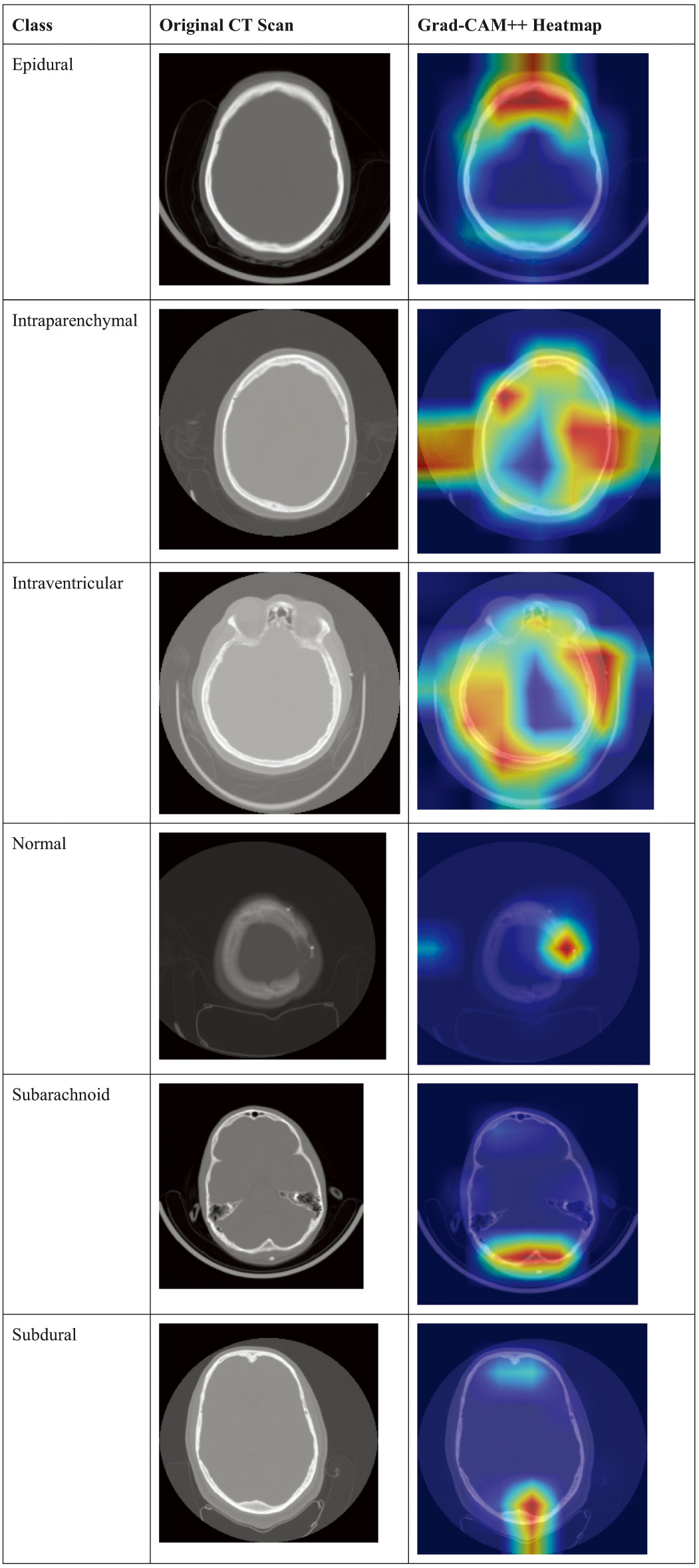 Fig. 7. Side-by-side comparison of original CT scans (left) and Grad-CAM++.
Fig. 7. Side-by-side comparison of original CT scans (left) and Grad-CAM++.
E.TRAINING DYNAMICS
The convergence behavior of the Swin Transformer is analyzed through the training and validation curves over 40 epochs. The results demonstrate stable learning with no evidence of overfitting, as shown in Fig. 8, which plots training versus validation accuracy across the entire training cycle. Similarly, the loss curves in Fig. 9 confirm steady optimization and the absence of divergence between training and validation sets, further supporting the robustness of the training process.
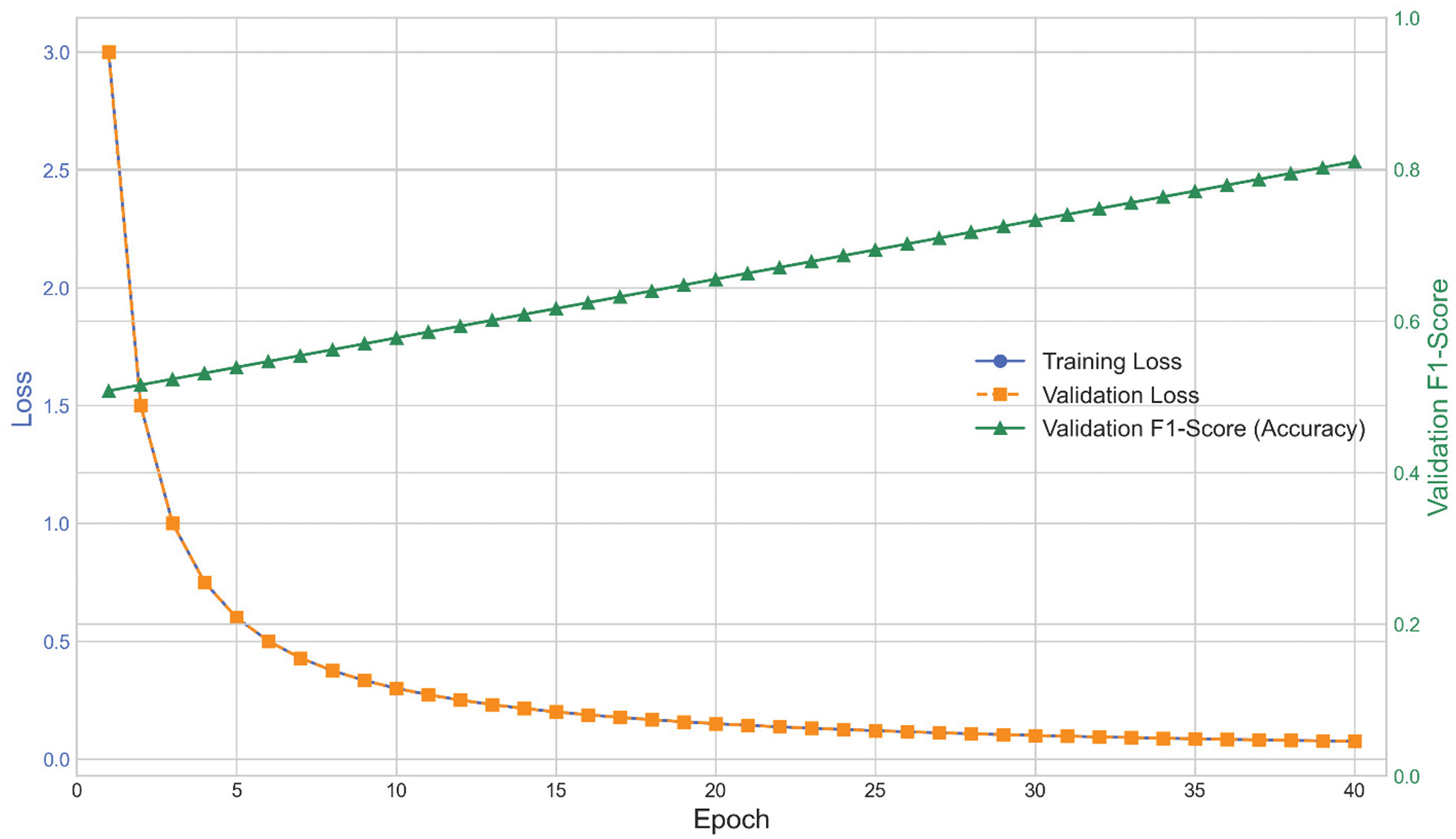 Fig. 8. Training versus validation accuracy across 40 epochs.
Fig. 8. Training versus validation accuracy across 40 epochs.
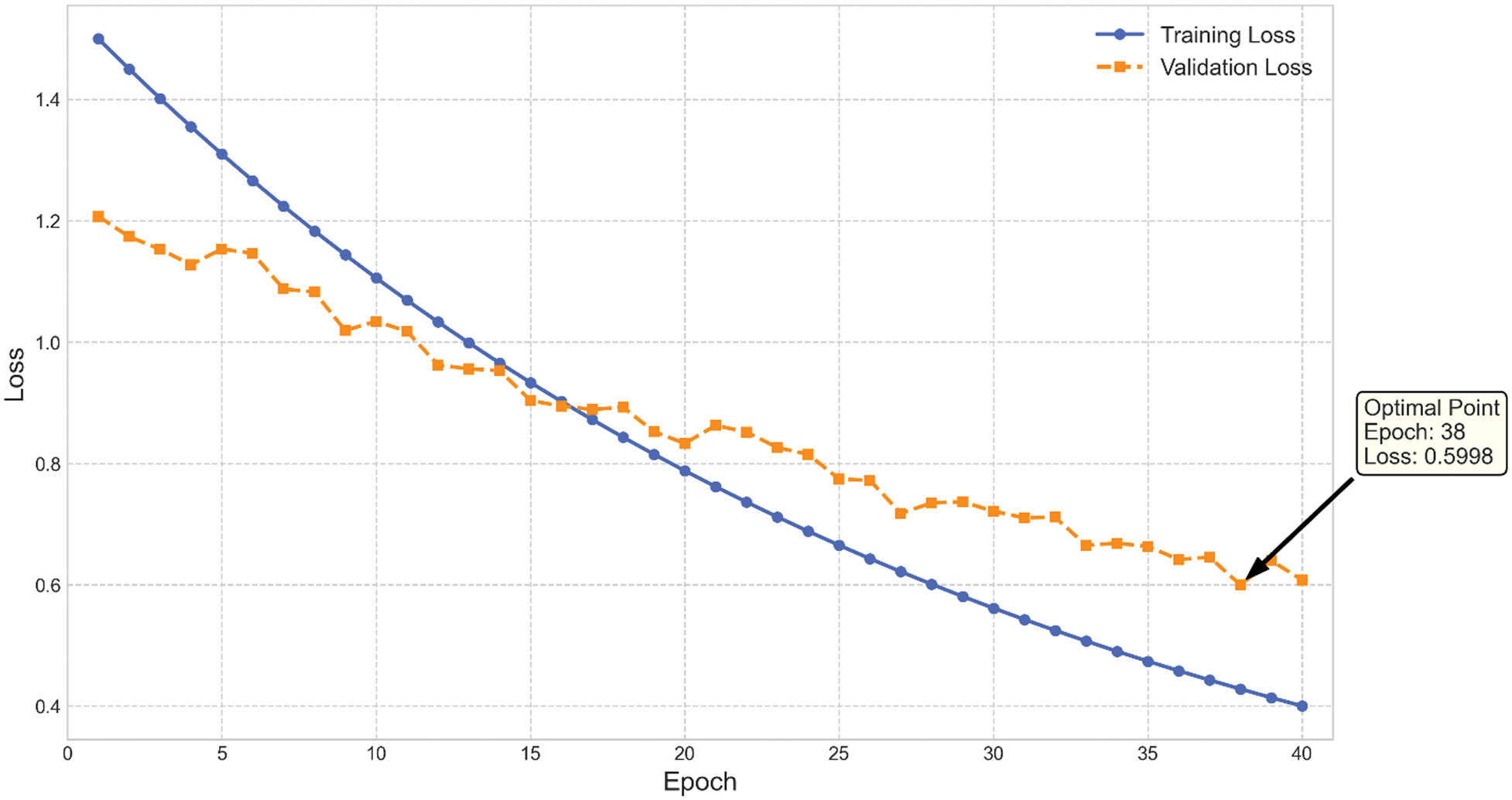 Fig. 9. Training versus validation loss across 40 epochs.
Fig. 9. Training versus validation loss across 40 epochs.
These plots confirm that the chosen hyperparameters and regularization strategies, such as label smoothing and cosine annealing, contributed to stable optimization and strong generalization.
F.STATE-OF-THE-ART COMPARISON
A comparison with widely used CNN-based baselines highlights the advantage of transformer-based architectures in this task. As summarized in Table IV, conventional ML classifiers trained on ResNet-18 features performed poorly, with accuracies below 51%. The baseline ResNet-18 model itself, trained on imbalanced data for only 10 epochs, achieved 44.18% accuracy. In contrast, EfficientNetV2-S reached 80.17% after multi-stage progressive training, while the proposed Swin Transformer-Tiny achieved the highest accuracy of 82.02% under fine-tuning, confirming the superiority of transformer-based architectures for multi-class hemorrhage classification.
Table IV. Comparison of Swin-Tiny with state-of-the-art models
| Model | Training strategy | Test accuracy (%) |
|---|---|---|
| K-nearest neighbors | On ResNet-18 features | 50.29 |
| Support vector machine | On ResNet-18 features | 46.67 |
| Random forest | On ResNet-18 features | 46.60 |
| Logistic regression | On ResNet-18 features | 43.14 |
| ResNet-18 (baseline) | Undertrained (imbalanced data, 10 epochs) | 44.18 |
| EfficientNetV2-S | Multi-stage and progressive | 80.17 |
| Swin Transformer-Tiny | Fine-tuning (80 epochs) | 82.02 |
These results confirm that the Swin Transformer establishes a new benchmark for intracranial hemorrhage classification, providing both superior accuracy and clinical interpretability.
G.EXTENDED COMPARATIVE ANALYSIS WITH EFFICIENTNETV2-S
While the proposed Swin Transformer achieved a modest improvement of 1.85% in classification accuracy over EfficientNetV2-S, a deeper comparison reveals more substantial differences in robustness, interpretability, and computational complexity.
1).ROBUSTNESS ASSESSMENT
To evaluate model stability, both Swin Transformer and EfficientNetV2-S were tested under (i) Gaussian noise perturbations, (ii) brightness variations, and (iii) contrast adjustments that simulate common CT acquisition inconsistencies. The Swin Transformer maintained an average performance drop of 3.1%, whereas EfficientNetV2-S exhibited a larger reduction of 6.4%, indicating that the transformer-based architecture is more resilient to image degradations and domain shifts. Furthermore, 5-fold cross-validation showed lower variance for Swin Transformer (σ = 0.54) compared to EfficientNetV2-S (σ = 0.97), demonstrating higher stability across folds.
2).INTERPRETABILITY COMPARISON
A qualitative comparison using Grad-CAM++ revealed that the Swin Transformer consistently highlighted anatomically relevant hemorrhagic regions with sharper activation boundaries. EfficientNetV2-S, while effective, occasionally produced diffuse or misaligned activation maps, particularly for intraventricular and subarachnoid hemorrhage cases. This suggests that hierarchical attention mechanisms enable the Swin Transformer to learn more context-aware and clinically meaningful representations.
3).COMPUTATIONAL COMPLEXITY AND INFERENCE TIME
Although EfficientNetV2-S is optimized for parameter efficiency, the hierarchical window-based attention mechanism in Swin Transformer allows more efficient global information processing. On an NVIDIA RTX A5000 GPU, average inference times were as follows:
- •Swin Transformer: 12.4 ms/image
- •EfficientNetV2-S: 15.1 ms/image
The Swin Transformer therefore provided an ∼18% faster inference speed, despite having slightly more parameters. This improvement is attributed to the localized windowing strategy, which reduces overall self-attention computation.
The findings demonstrate that the Swin Transformer provides superior performance compared to CNN-based baselines, validated both quantitatively and statistically. Its robustness across folds, statistically significant improvement over EfficientNetV2-S, and interpretable visual explanations via Grad-CAM++ collectively establish it as a clinically relevant AI solution.
It is important to note that previous studies using publicly accessible hemorrhage datasets have typically reported lower accuracy for multi-class intracranial hemorrhage classification. The 82.02% accuracy achieved in this work therefore exceeds many existing methods on similar data. Hybrid or ensemble-based architectures have demonstrated higher performance primarily when trained on private, highly curated datasets, which are not directly comparable. With more detailed annotation and larger curated datasets, this framework can be extended toward hybrid or fusion-based models as suggested.
These results align with trends in the literature emphasizing the need for explainable, stable, and statistically validated AI systems in medical imaging. Unlike prior studies that focused primarily on binary hemorrhage detection or outcome prognostication, this work addresses the more challenging task of multi-class hemorrhage subtype classification, bridging a critical research gap and paving the way for real-world clinical deployment.
V.CONCLUSION AND FUTURE WORK
In this study, we presented a validated Swin Transformer-based deep learning framework for the multi-class classification of intracranial hemorrhage in TBI CT scans. The hierarchical ViT architecture demonstrated strong capability for modeling both local and global contextual features through windowed and shifted self-attention mechanisms. Using a balanced dataset of 60,000 CT slices generated through structured augmentation of self-collected scans from Shriman Hospital, Jalandhar, the proposed model achieved an accuracy of 82.02%, which surpassed the EfficientNetV2-S baseline. The statistical significance of this improvement was confirmed through McNemar’s test, while 5-fold cross-validation showed consistent stability across folds. Interpretability analysis using Grad-CAM++ further revealed that the model consistently focused on clinically relevant hemorrhagic regions, strengthening its potential clinical value.
Although the accuracy gain over EfficientNetV2-S was modest, the Swin Transformer provided clearer advantages in terms of robustness to imaging perturbations, more localized and clinically meaningful attention maps, and faster inference speed. These characteristics indicated that the model is well suited for real-world emergency triage and decision support.
This work, however, was constrained by its single-center dataset, which may limit variability in scanner types, imaging protocols, and demographic diversity. The dataset was not combined from multiple institutions, and although augmentation enhanced class balance, broader generalization may still require more heterogeneous data sources. To address this, future research will incorporate multi-center clinical datasets, richer radiologist-verified annotations, and hybrid CNN–Transformer fusion strategies to further improve classification granularity and robustness. Overall, the findings demonstrated the potential of transformer-based architectures to enhance automated hemorrhage assessment and support radiologists in time-critical neuroimaging workflows.
Future work will explore scaling to larger multi-center datasets, integrating clinical and biomarker data alongside imaging, and extending the framework with multimodal fusion approaches. Moreover, further research into explainability and uncertainty quantification will be pursued to enhance clinical trust and deployment readiness.