I.INTRODUCTION
When the fluid-filled spaces in the fetal brain are larger than normal, the condition is known as fetal ventriculomegaly (VM). Doctors usually classify how large these spaces are by measuring their width. If the width is between 10 and 12 millimeters, it is considered mild. Moderate VM is when the width is 13 to 15 millimeters, and anything wider than 15 millimeters is severe [1,2]. In cases of mild VM, the likelihood of the infant experiencing developmental delays is significantly lower. In fact, research shows that around 7.9% of babies experience neurodevelopmental delay in case of mild VM which can be there in the general population [3].
In another comprehensive review, it is stated that mild isolated fetal VM is associated with a significantly more favorable prognosis compared to more severe cases. In the case of severe VM, there are 4.24 relative risks for neurodevelopmental delay, 4.46 for intrauterine fetal demise, and 6.02 for infant mortality in comparison with mild VM [4,5]. It is challenging to predict how VM will affect a baby’s development and because of this, it is very difficult for the doctors and healthcare providers to make decisions and to counsel parents. Currently, doctors focus on the size of the brain’s fluid-filled spaces (ventricles) and qualitative assessments. However, these methods are not always accurate in forecasting whether the baby will have developmental issues later on. Recently, AI and deep learning tools have provided significant steps in the analysis of medical images, and based on those, predictions can be made. This represents a promising approach to improving the accuracy of VM diagnosis and management [6–10]. The prevalence of impaired fetal brain development is approximately 3 per 1,000 pregnancies [11]. VM is the term used when a doctor or sonographer detects enlargement of the fetal brain’s lateral ventricles. With a prevalence of about 7.03 per 10,000 births, VM is one of the more recently identified abnormalities in the development of the fetal central nervous system [12].
Deep learning provides a strong approach for the analysis of medical images [13,14]. Architectures like Convolutional Neural Network (CNN) and attention-based architectures are effective for a vast range of applications, including tumor segmentation, organ classification, and the detection of diseases [15–17]. DenseNet and EfficientNet architectures have shown great feature extraction capabilities, while attention mechanisms like Squeeze-and-Excitation (SE) and Convolutional Block Attention Module (CBAM) have been utilized for enhancing the importance of features by emphasizing prominent regions. Though many innovations prevail, the challenges for the aim of balancing the computational efficiency with the level of accuracy continue for the purposes of the clinical practice, especially for the health settings with limited resources.
To address the gaps, we propose a new CAAF (Cross-Attention and Adaptive Fusion) framework. We employ lightweight convolutional backbones and channel-spatial attention and cross-attention fusion layers for optimal discriminative capability and reduced computational complexity, respectively. Comparing thoroughly with baseline architectures (viz., customized CNN, DenseNet, DenseNet with SE-CBAM, EfficientNet with SE-CBAM, and DenseNet–EfficientNet fusion), our proposed approach demonstrates excellent ability for VM detection from fetal ultrasound images.
The remainder of the paper is schematized as follows. The literature review summary is defined in Section II. Section III deliberates the methodology, which includes the dataset, proposed framework (both Stage 1 and Stage 2), and algorithms. Section IV reports experimental outcomes and comparative analysis. Section V discusses findings and limitations, and Section VI concludes the paper.
II.LITERATURE REVIEW
The detection of fetal VM is an issue that has captured significant research attention over the past few years. Moreover, the emphasis of such recent research is also being directed toward deep learning-based segmentation, classification, as well as hybrid approaches to improve the reliability of the detection process. Although most of the earlier works have been based on handcrafted features as well as traditional machine learning approaches, recent research is also being directed toward CNNs, attention-based approaches, as well as multimodal approaches. In this context, a summary of the recent advancements by the research community is provided for fetal VM detection in four categories: segmentation-based detection approaches, classification-based approaches, attention-based approaches, as well as explainable approaches. In such a changing paradigm, we proposed our novel framework – CAAF.
The researchers Chen et al. employed machine learning techniques on prenatal Magnetic Resonance Imaging (MRI) data to predict post-delivery ventricular irregularities in fetuses with isolated VM. Their model provides analysis of intracranial structures and brain parenchyma radiomics surrounding the occipital horn of the lateral ventricle. They demonstrated the potential of radiomics in prognostication. While radiomics models show promise, their generalizability across different imaging protocols and populations remains a challenge. They suggested to use multimodal data which integrates genetic, imaging, and clinical data to develop the comprehensive prognostic models [18].
In the recent study done by Vahedifard et al., they focused on an automated strategy using a deep learning model to estimate the lateral ventricle linearly in fetal brain MRI. They categorized them into two classes: normal and VM. They claim that the performance of deep learning models is heavily reliant on the quality and quantity of annotated data. Transfer learning approaches can be used to leverage pretrained models and adapt them to specific datasets, which may improve model performance with limited data [19].
Attention-Guided CNN (AG-CNN) for maternal–fetal ultrasound image analysis was introduced by Muna Al-Razgan et al. to achieve high accuracy rates. Their attention mechanisms enabled the model to focus on diagnostically pertinent regions to improve performance. According to the authors, deep learning models that utilize attention mechanisms require careful interpretation to ensure that the identified regions align with clinical expectations. For future work, they suggested developing methods to augment the interpretability of deep learning models to ensure their decisions are transparent and clinically relevant [20].
In the research, Li et al. proposed the recurrent convolutional network to forecast the progression of mild VM based on the initial imaging features. The researchers also considered clinical factors in their proposed model to forecast whether the mild cases would resolve, remain stable, or progress to moderate–severe VM. The proposed method is restricted to predicting short-term radiological progression rather than long-term neurodevelopmental outcome [21].
In the research paper by Zhigao Cai et al., the researchers developed a new framework for the dissection of fetal brain MRI images using attention mechanisms, depth-wise distinguishable convolution, and a cascade network. In the paper, the researchers were able to address the problems of motion correction of the fetal brain for precise brain extraction. The researchers observed that the use of attention-based models may be computationally rigorous and require a lot of computational power. The researchers also recommended the development of lightweight attention-based models for real-time processing to support real-time decision-making [22].
In the study by Mohammad Asif Hasan et al., the authors used multi-head attention (MHA) mechanisms to predict the fetal brain’s gestational age using diffusion tensor imaging (DTI). The effectiveness of the attention mechanism in handling sequential data was clearly shown in this study, and the accuracy of the predictions was enhanced. The authors stated that the effective training of the attention mechanism requires access to large and high-quality datasets, which may not be available in all cases. The authors further suggested the potential use of the attention mechanism in various imaging modalities to boost the accuracy of the diagnostic tools [23].
We also came across a study done by Gopikrishna and his team in which they used U-Net and DeepLab V3+ for different segmentation tasks. They concluded that DeepLab V3+ is more accurate for segmentation tasks, whereas U-Net is more accurate for classification tasks. However, one thing that we learned from this study is that none of the models is best for all tasks; hence, we need a combination of models for optimal performance [24].
Another important study was done by Zhu and his team in which they proposed a DenseNet-based model integrated with the Schmidt Neural Network for brain disease classification. They were successful in attaining high accuracy with optimal overfitting. This study indicates that DenseNet is a reliable model for medical image classification problems [25].
In another study done by Sengodan, DenseNet was used for histopathology image classification. In this study, it was concluded that EfficientNet integrated with attention models like CBAM reaches an accuracy level of nearly 99%. This study is important because it indicates that attention models are helpful in training models that focus on key parts of the image for accurate classification [26].
Gopikrishna et al. employed DenseNet in association with segmentation and texture-based feature engineering for brain tumor classification. Their model recorded an accuracy of 97%. The authors’ research established the efficacy of integrating deep learning with traditional feature engineering to improve the accuracy of brain tumor classification [27].
The paper published by Fakheri et al. proposed an effective deep learning model based on DenseNet for brain tumor classification. The model achieved an accuracy of 96.96%, along with high sensitivity (98.84%), specificity (93.43%), and precision (96.59%), outperforming traditional models such as Backpropagation (BP), U-Net, and Region-Based Convolutional Neural Network (RCNN). The study demonstrated the effectiveness of integrating segmentation, feature engineering, and deep learning techniques to enhance classification performance. Furthermore, the proposed model serves as a valuable tool for radiologists, enabling automated, reliable, and efficient classification of brain tumors [28].
Shirsat and team applied CNN, separable CNNs, and Xception for fetal brain anomaly detection in ultrasound. They pointed out that while anomaly detection was feasible, distinguishing between mild and moderate VM remained a significant challenge [29].
Table I. Summary of literature review
| Study (year) | Modality | Methodology | Task | Key contributions | Limitations/gaps |
|---|---|---|---|---|---|
| Chen et al. (2024) | Prenatal MRI | Radiomics + ML | Prognosis of isolated VM | Demonstrated predictive value of parenchymal and ventricular radiomics around occipital horn | Limited generalizability across scanners; requires MRI; lacks ultrasound applicability |
| Vahedifard et al. (2023) | Fetal MRI | CNN-based linear measurement | Binary VM detection | Automated and consistent ventricle measurement outperforming manual methods | Binary classification only; dependent on annotated MRI data |
| Al-Razgan et al. (2024) | Ultrasound | AG-CNN (Attention-Guided CNN) | Fetal anomaly detection | Attention improves localization of diagnostically relevant regions | Limited interpretability validation; no severity grading |
| Li et al. (2021) | MRI | Recurrent CNN + clinical data | VM progression prediction | Integrated temporal imaging and clinical factors | Short-term prediction only; no ultrasound validation |
| Cai and Zhao (2024) | Fetal MRI | Cascade CNN + attention | Brain segmentation | High-precision fetal brain extraction with motion robustness | Computationally expensive; not suited for real-time or low-resource settings |
| Hasan et al. (2024) | DTI (MRI) | Multi-Head Attention + Xception | Gestational age prediction | Demonstrated effectiveness of attention for sequential neuroimaging | Requires large datasets; modality-specific |
| Gopikrishna et al. (2024) | MRI | U-Net vs DeepLabV3+ | VM segmentation | Showed trade-off between segmentation accuracy and classification performance | No unified architecture for both tasks |
| Zhu et al. (2022) | MRI | DenseNet + SNN | Brain disease classification | High accuracy with reduced overfitting | Not applied to fetal ultrasound or VM |
| Sengodan (2025) | Histopathology | EfficientNet + CBAM | Image classification | Highlighted power of attention-guided EfficientNet | Domain mismatch with fetal imaging |
| Gopikrishna et al. (2025) | MRI | DenseNet + explainability | VM classification | Strong interpretability via explainable ML | MRI-only; lacks ultrasound validation |
| Fakheri et al. (2024) | MRI | DenseNet-based framework | Brain tumor classification | High sensitivity and specificity; clinical viability | Task-specific; not transferable to VM grading |
| Shirsat et al. (2025) | Ultrasound | CNN, Separable CNN, Xception | Fetal anomaly detection | Demonstrated feasibility on ultrasound | Difficulty distinguishing mild vs moderate VM |
By reviewing these studies, we identified the following insights:
- •Several models are available, among which DenseNet and EfficientNet serve as strong backbone models.
- •Integrating attention mechanisms, such as SE and CBAM, enhances performance by directing the network toward key features.
- •Distinguishing between different categories of VM remains a significant challenge, even with advanced models.
To the best of our knowledge, no previous work has explored a multi-backbone, attention-guided fusion approach for this task. Our research addresses this gap by systematically benchmarking several models and introducing the novel CAAF architecture. In our proposed CAAF framework, we combine DenseNet and EfficientNet with SE + CBAM and a novel cross-attention fusion strategy where features are extracted by modern CNNs, attention mechanisms sharpen clinical focus, and cross-attention enhances joint representation learning. A comprehensive summary of the reviewed studies, including their methodologies, key contributions, and limitations, is presented in Table I.
III.METHODOLOGY
A.DATASET
The dataset used in this study was obtained from the publicly available Fetal Brain Abnormalities Ultrasound collection hosted on Roboflow Universe (Hritwik Trivedi, 2023; CC BY 4.0 license). It comprises anonymized ultrasound scans of fetal brains, classified into four clinically validated categories: Normal, Mild VM, Moderate VM, and Severe VM.
The dataset has been divided into training, validation, and testing sets, as shown in Table II. The stratified sampling technique has been applied to ensure the class balance in the datasets. The images have been resized to a size of 380 × 380 pixels, matching the input size required for the EfficientNetB4 model, without compromising the essential anatomical features of the images.
Table II. Dataset distribution
| Class | Train | Validation | Test |
|---|---|---|---|
| Mild ventriculomegaly | 195 | 27 | 24 |
| Moderate ventriculomegaly | 258 | 30 | 26 |
| Normal | 227 | 31 | 24 |
| Severe ventriculomegaly | 94 | 13 | 19 |
Table III. Preprocessing and augmentation – summary of image preprocessing and data augmentation techniques applied during training. Standardized normalization and controlled geometric augmentations were employed to enhance model generalization while preserving clinically relevant anatomical structures in fetal brain ultrasound images
| Parameter | Value | Rationale/code reference |
|---|---|---|
| Image resolution | 380 × 380 pixels | Matches IMG_SIZE and EfficientNet-B4 input size. |
| Normalization | EfficientNet preprocessing | Standardized to [−1, 1] range. |
| Data augmentation | RandomFlip(“horizontal”), RandomRotation(0.05), RandomZoom (0.05) | Applied via tf.keras.Sequential in make_dataset. |
Table IV. Training configurations - training hyperparameters and optimization settings used during the two-stage training procedure of the CAAF model. The staged freezing and fine-tuning strategy balances convergence stability and feature adaptability while mitigating overfitting in limited medical datasets
| Parameter | Stage 1 (initial) | Stage 2 (fine-tuning) |
|---|---|---|
| Optimizer | Adam | Adam |
| Learning rate | 1 × 10−4 | 1 × 10−5 (decay for fine-tuning) |
| Batch size | 8 | 8 |
| Backbone status | Frozen | Top layers unfrozen (DenseNet121 last 50, EfficientNet-B4 last 40) |
| Loss function | Sparse categorical cross-entropy (SCCE) | Multi-loss: SCCE (classification) + alignment loss (mask refiner) |
| Loss weights | Classification: 1.0 | Classification: 1.0, prior refiner: 0.3 |
| Class balancing | Class weights (balanced) | Class balancing was applied only in Stage 1 to correct dataset imbalance during representation learning, while Stage 2 focuses on refining spatial priors and therefore omits class-weighted optimization. |
In order to validate the generalization capability of the framework, the framework was further validated on the Zenodo Trans-Ventricular dataset, comprising 597 images [32].
B.BASELINE MODELS
For the purpose of developing a robust performance reference point for the proposed framework, a rigorous performance evaluation of different baseline approaches was conducted. These approaches represent different design strategies for medical image analysis. For the purpose of a fair performance comparison of the different approaches, all the approaches under consideration were trained under the same experimental conditions. This implies that all the models under consideration have been trained using the same dataset split, image size, preprocessing, and optimizer configuration. In addition, the issue of class imbalance has been addressed for all the approaches under consideration. A uniform weighting strategy has been adopted for addressing the issue of class imbalance.
1).CUSTOM CNN
A lightweight CNN has been implemented, comprising three convolutional layers, batch normalization, and dropout. Although the proposed lightweight CNN has shown poor performance in terms of accuracy (accuracy: 0.688), it has shown inadequate performance for the complex VM classification task. The confusion matrix for the lightweight custom CNN model, tested on the test set, has shown high inter-class confusion, particularly between mild and moderate VM. The confusion matrix of the custom CNN model is presented in Fig. 1, illustrating its classification performance across different classes.
2).DenseNET121
We fine-tuned a DenseNet121 model that was pretrained on the ImageNet database. This resulted in a significant improvement in accuracy to 0.849. This proved the importance of transfer learning in improving medical image classification tasks with less training data. Confusion matrix for the DenseNet121 model pretrained on ImageNet and fine-tuned for VM classification. The model appears to have better class separation compared to the custom CNN model. This is a clear example of the importance of transfer learning in fetal ultrasound image analysis. However, there is still some level of misclassification between consecutive levels of severity. The classification performance of DenseNet121 is depicted in Fig. 2 through its corresponding confusion matrix.
3).DenseNet + SE-CBAM
The DenseNet121 model was enhanced with channel-spatial recalibration by incorporating SE and CBAM modules. This resulted in a model that was able to achieve 0.957 in accuracy while maintaining consistency across all classes. This was enabled by the application of attention mechanisms to focus on areas of interest for diagnosis. Confusion matrix for DenseNet121 enhanced with SE and CBAM. The integration of channel-spatial attention helps to enhance consistency in classification across all levels of severity in VM. This is achieved by focusing on areas of interest in the ventricles for diagnosis. The confusion matrix of the proposed DenseNet model enhanced with SE attention and CBAM is shown in Fig. 3, highlighting improved class-wise predictions.
4).EfficientNet + SE-CBAM
EfficientNetB4 augmented with SE and CBAM modules experienced training collapse (accuracy =0.258), likely due to overparameterization relative to the dataset size. The confusion matrix for the EfficientNet-based model integrated with SE attention and CBAM is presented in Fig. 4.
5).DenseNet + EfficientNet GATED FUSION
Hybrid concatenation of feature embeddings from both DenseNet and EfficientNet achieved accuracy of 0.957 with high consistency and balanced precision-recall trade-offs. This demonstrated the value of multi-backbone approaches. The model achieves balanced precision and recall across classes by combining complementary features from two heterogeneous backbones; however, its fixed fusion strategy restricts adaptability to case-specific anatomical variations. The performance of the DenseNet–EfficientNet gated fusion model is illustrated in Fig. 5 using its confusion matrix.
C.PROPOSED METHODOLOGY – CAAF FOR PREDICTING VM IN FETAL BRAIN
1).ARCHITECTURE OVERVIEW
The proposed CAAF framework integrates dual CNN backbones – DenseNet121 and EfficientNetB4 – with lightweight attention and fusion mechanisms. The architecture is designed to achieve three primary goals:
- 1.Enhance feature expressiveness through cross-attention between two heterogeneous networks.
- 2.Enable adaptive feature fusion via gated weighting to select class-relevant features dynamically.
- 3.Maintain computational efficiency by employing lightweight 1 × 1 convolutional transformations and reduced projection dimensions.
CAAF operates in two stages:
- •Stage 1: Independent feature extraction by DenseNet121 and EfficientNetB4.
- •Stage 2: Bidirectional cross-attention followed by gated adaptive fusion (GAF), culminating in a fully connected classification head.
The architecture operates in two stages, as illustrated in Figs 6 and 7.
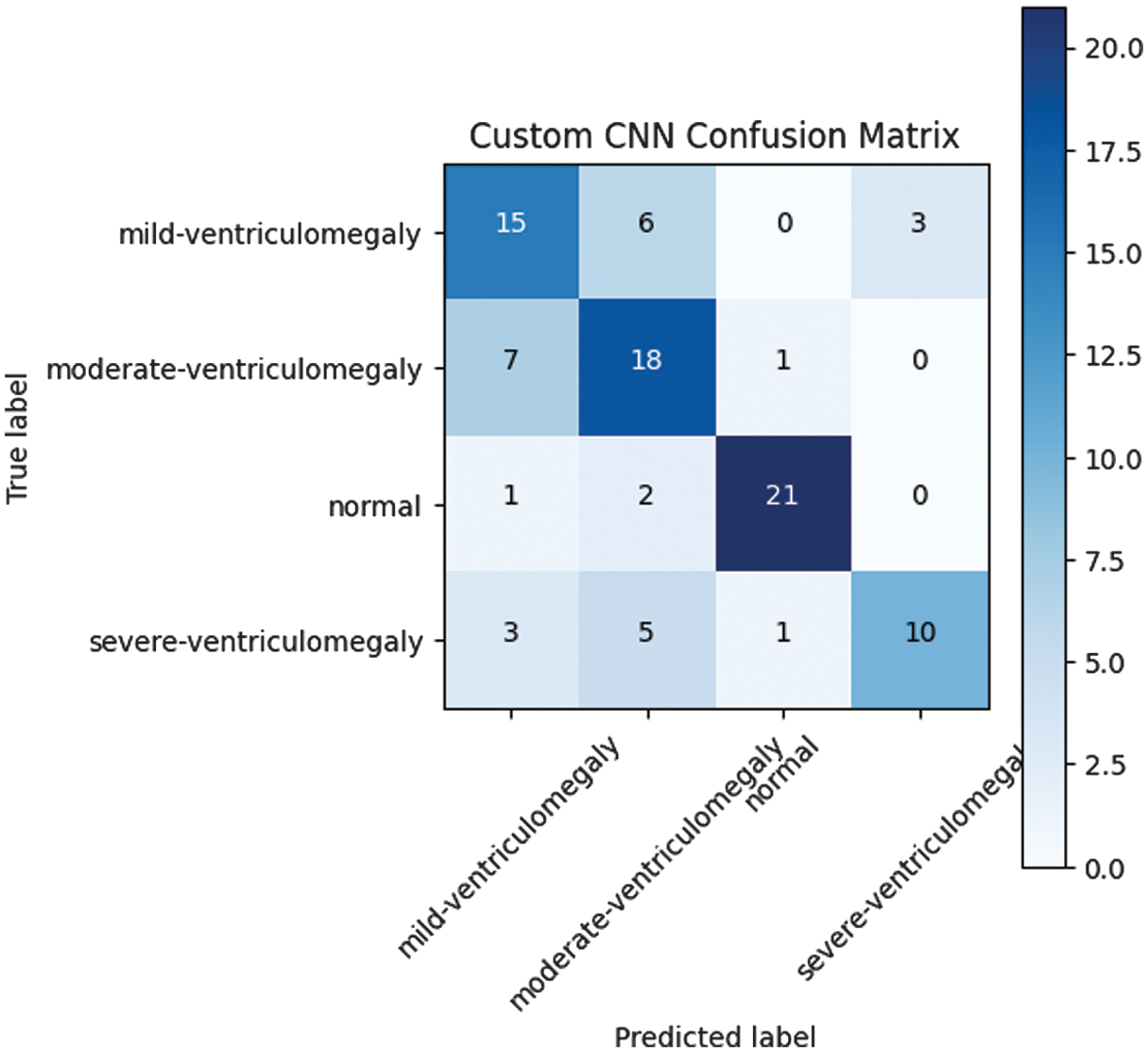 Fig. 1. Custom CNN confusion matrix.
Fig. 1. Custom CNN confusion matrix.
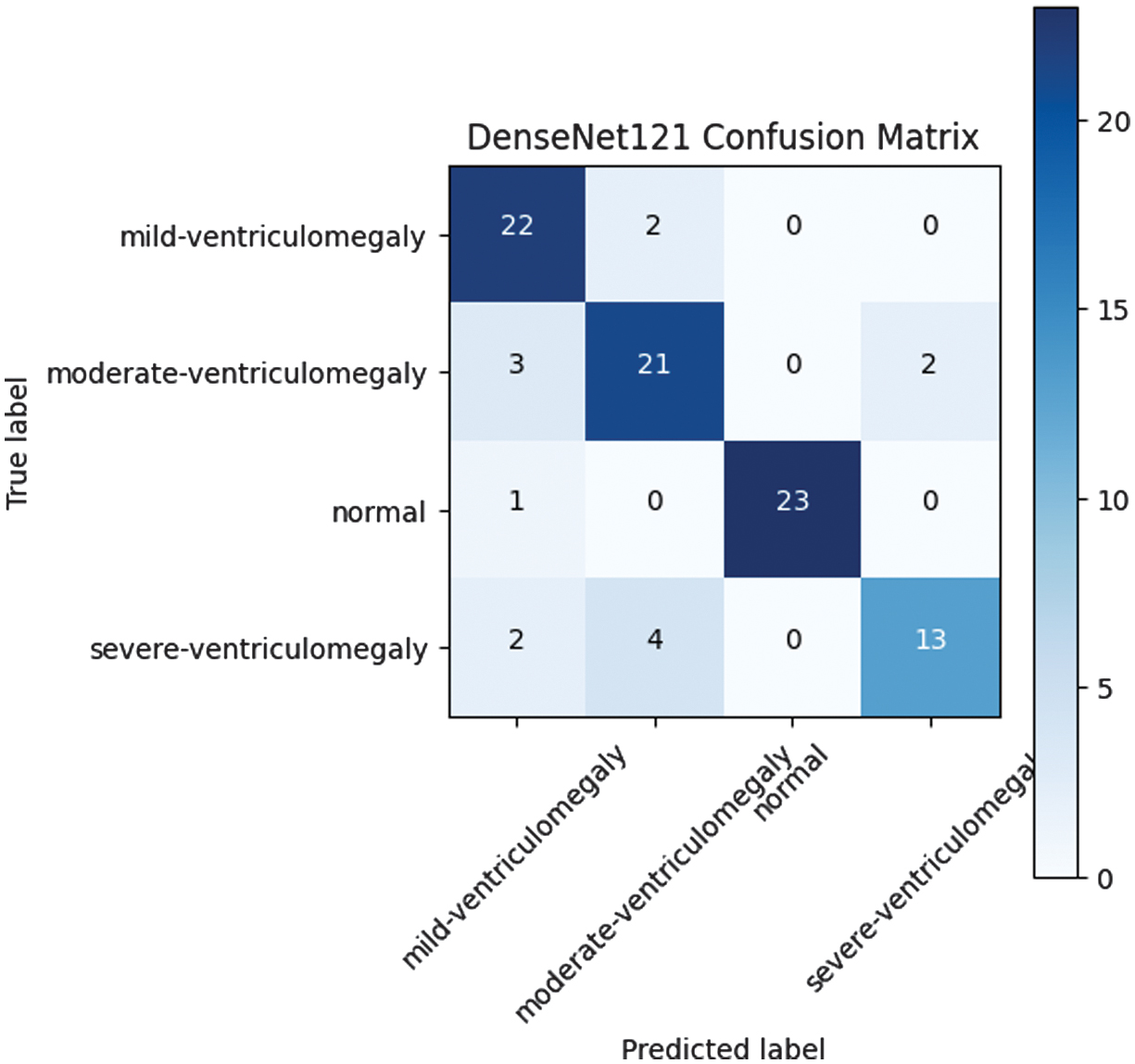 Fig. 2. DenseNet121 confusion matrix.
Fig. 2. DenseNet121 confusion matrix.
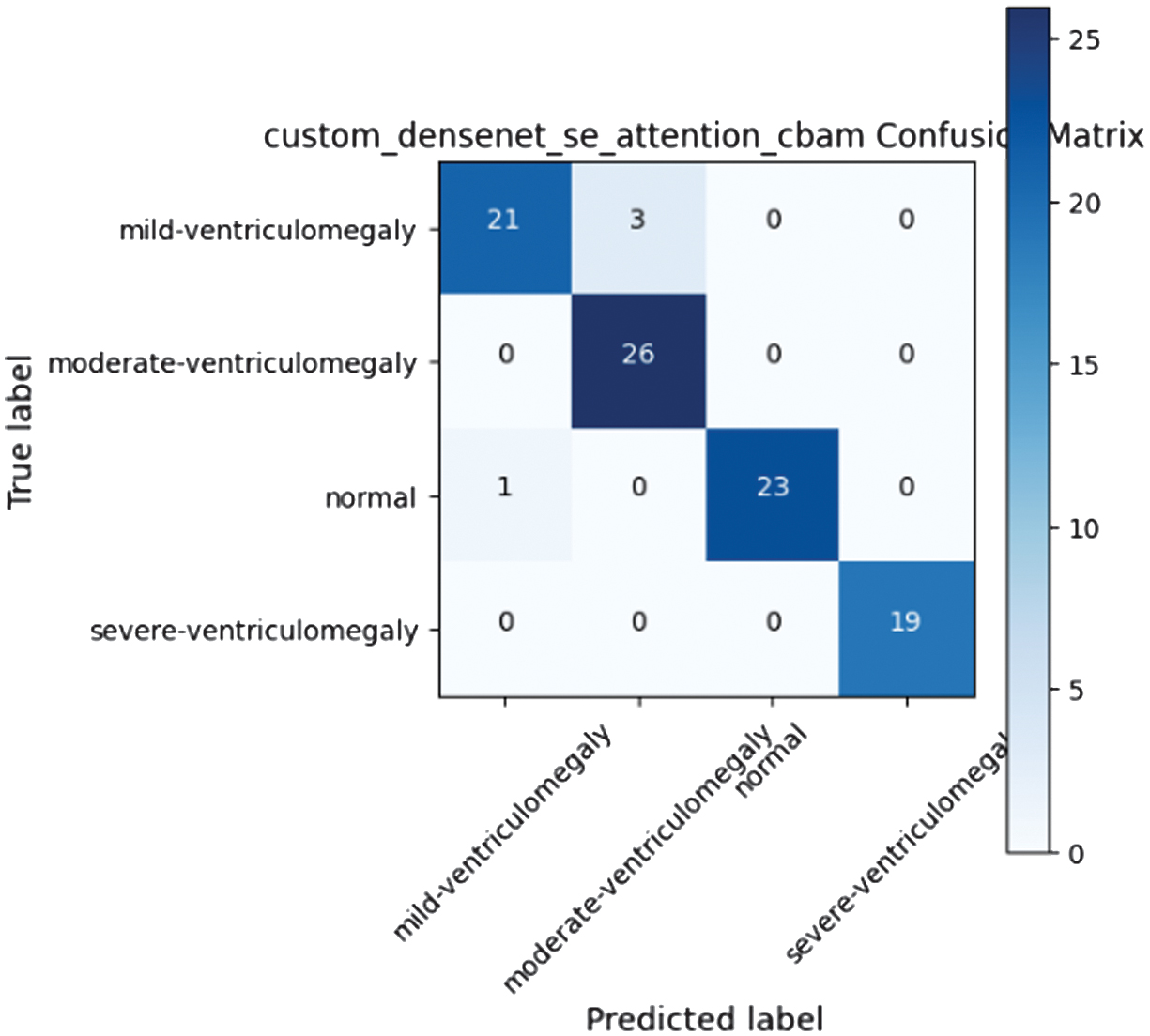 Fig. 3. Custom DenseNet SE attention CBAM confusion matrix.
Fig. 3. Custom DenseNet SE attention CBAM confusion matrix.
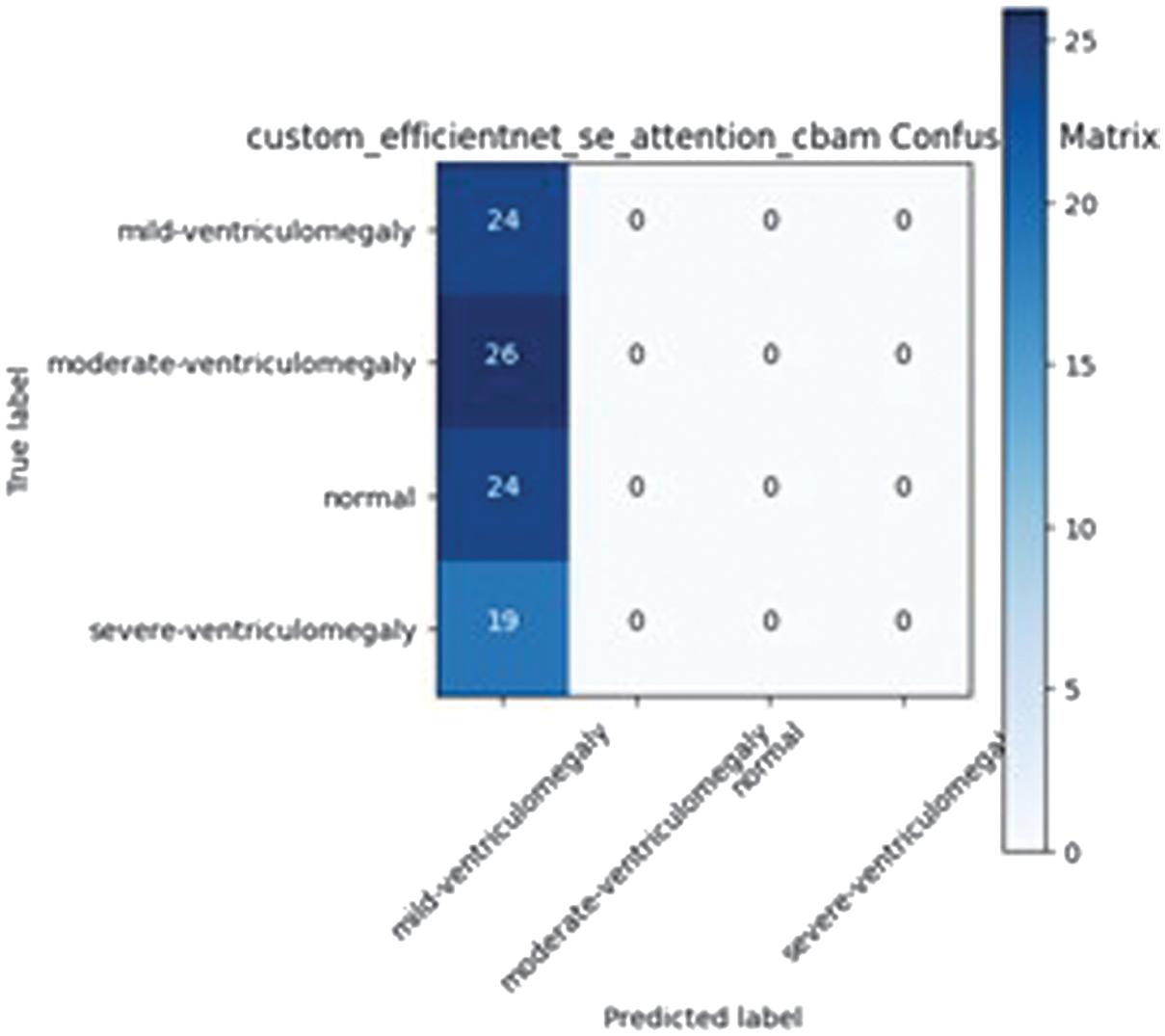 Fig. 4. Custom EfficientNet SE attention CBAM confusion matrix.
Fig. 4. Custom EfficientNet SE attention CBAM confusion matrix.
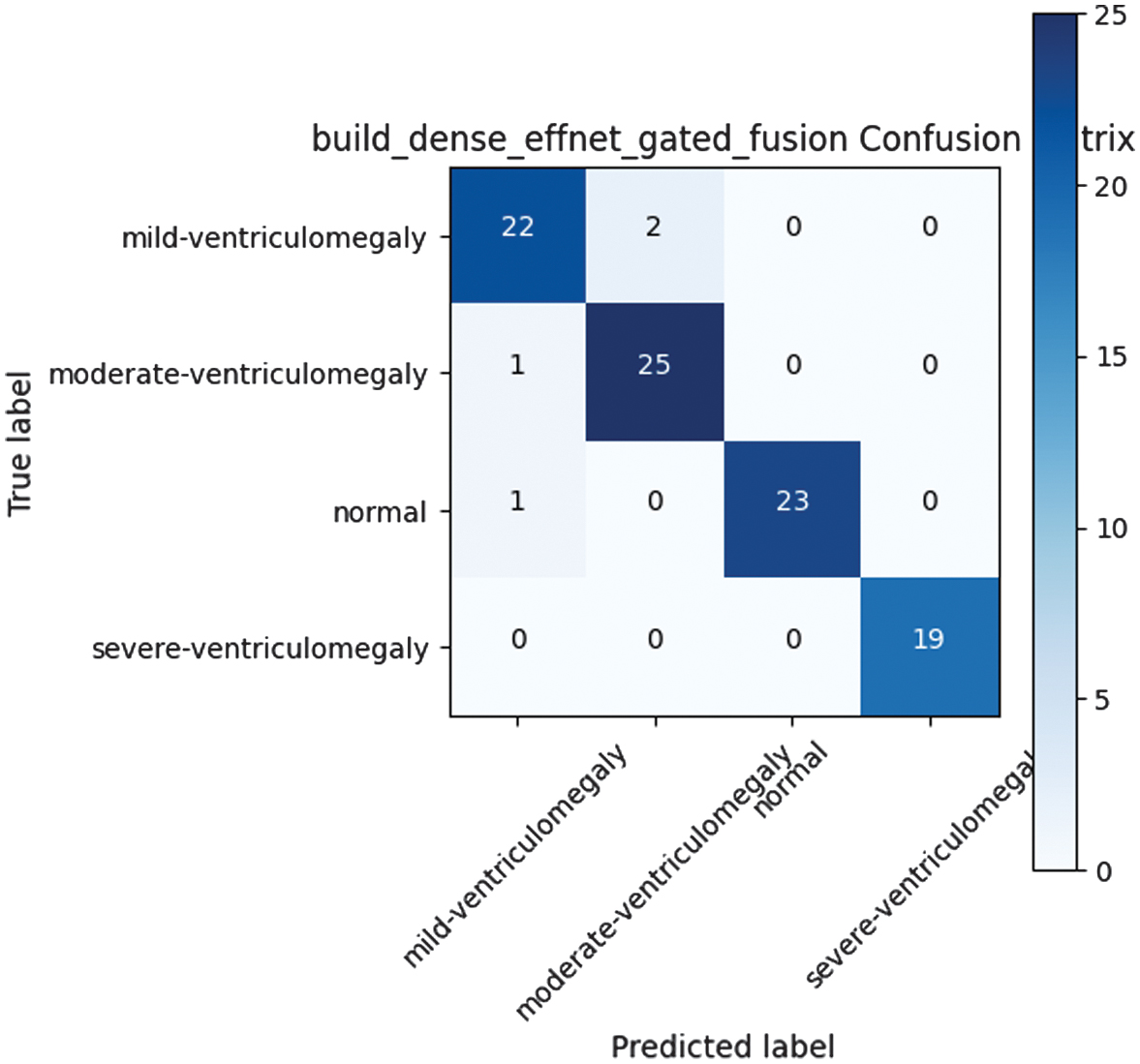 Fig. 5. DenseNet-EfficientNet gated fusion confusion matrix.
Fig. 5. DenseNet-EfficientNet gated fusion confusion matrix.
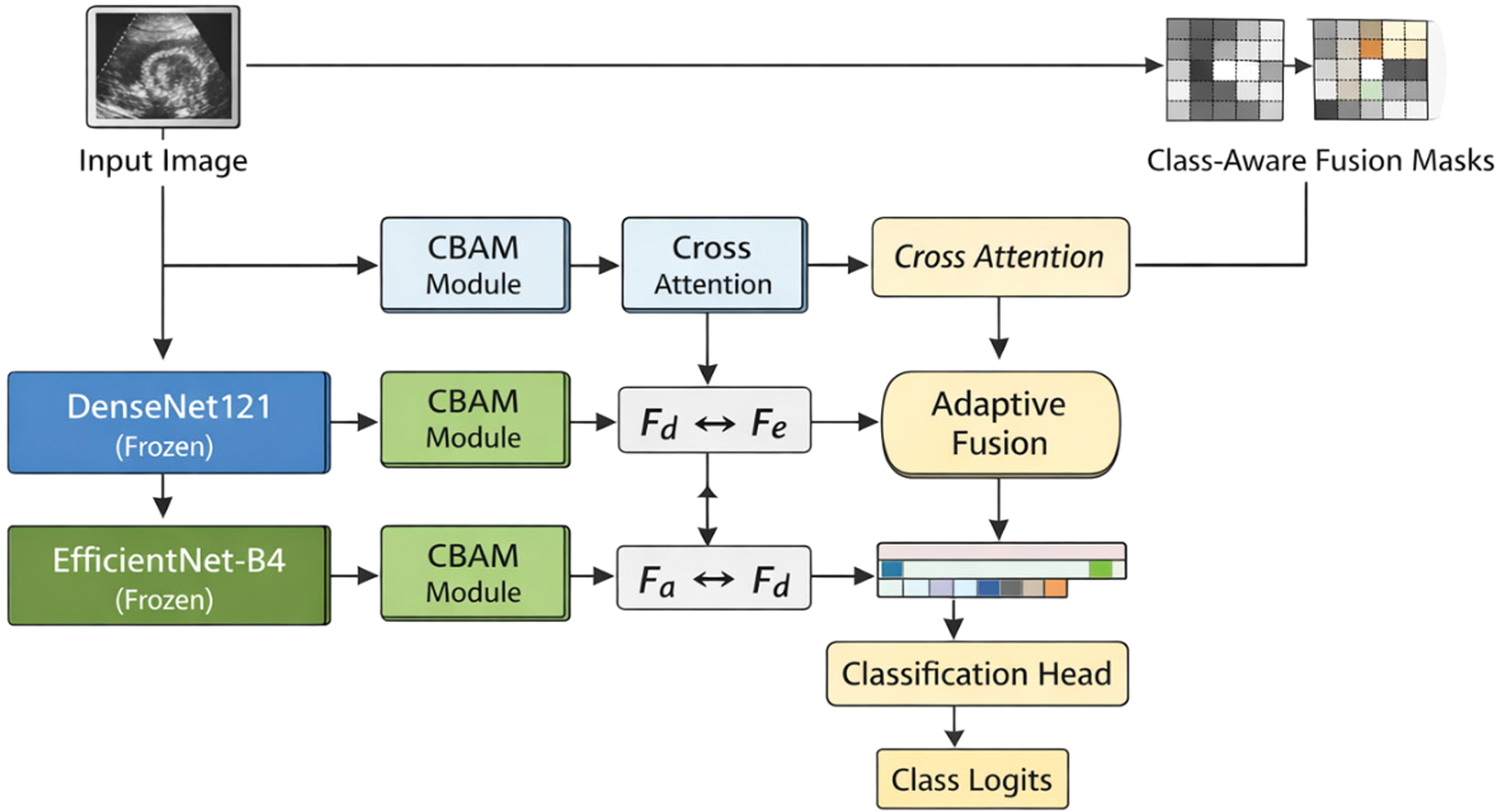 Fig. 6. Architecture of the proposed CAAF model (Stage 1).
Fig. 6. Architecture of the proposed CAAF model (Stage 1).
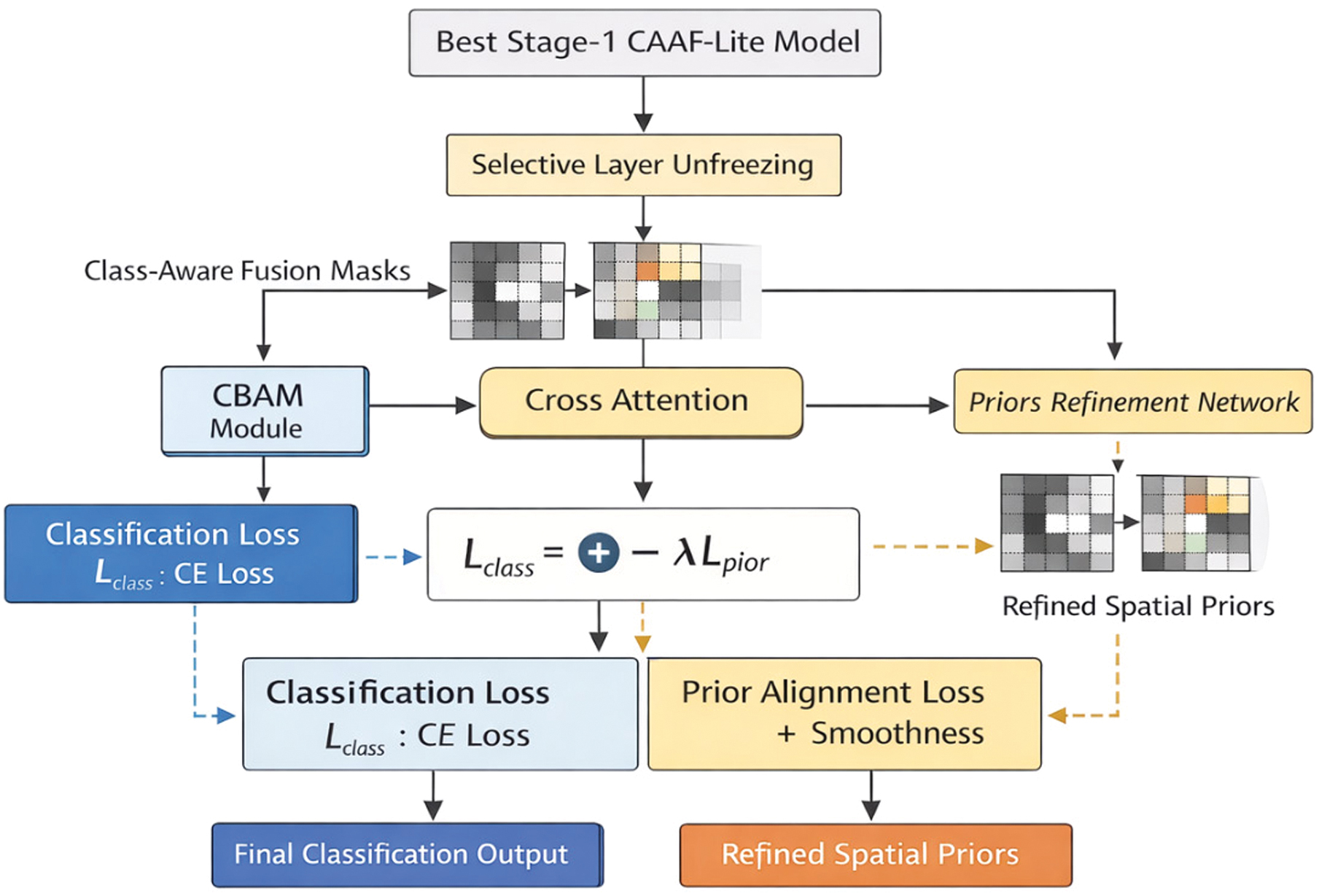 Fig. 7. Stage 2 prior-guided fine-tuning framework.
Fig. 7. Stage 2 prior-guided fine-tuning framework.
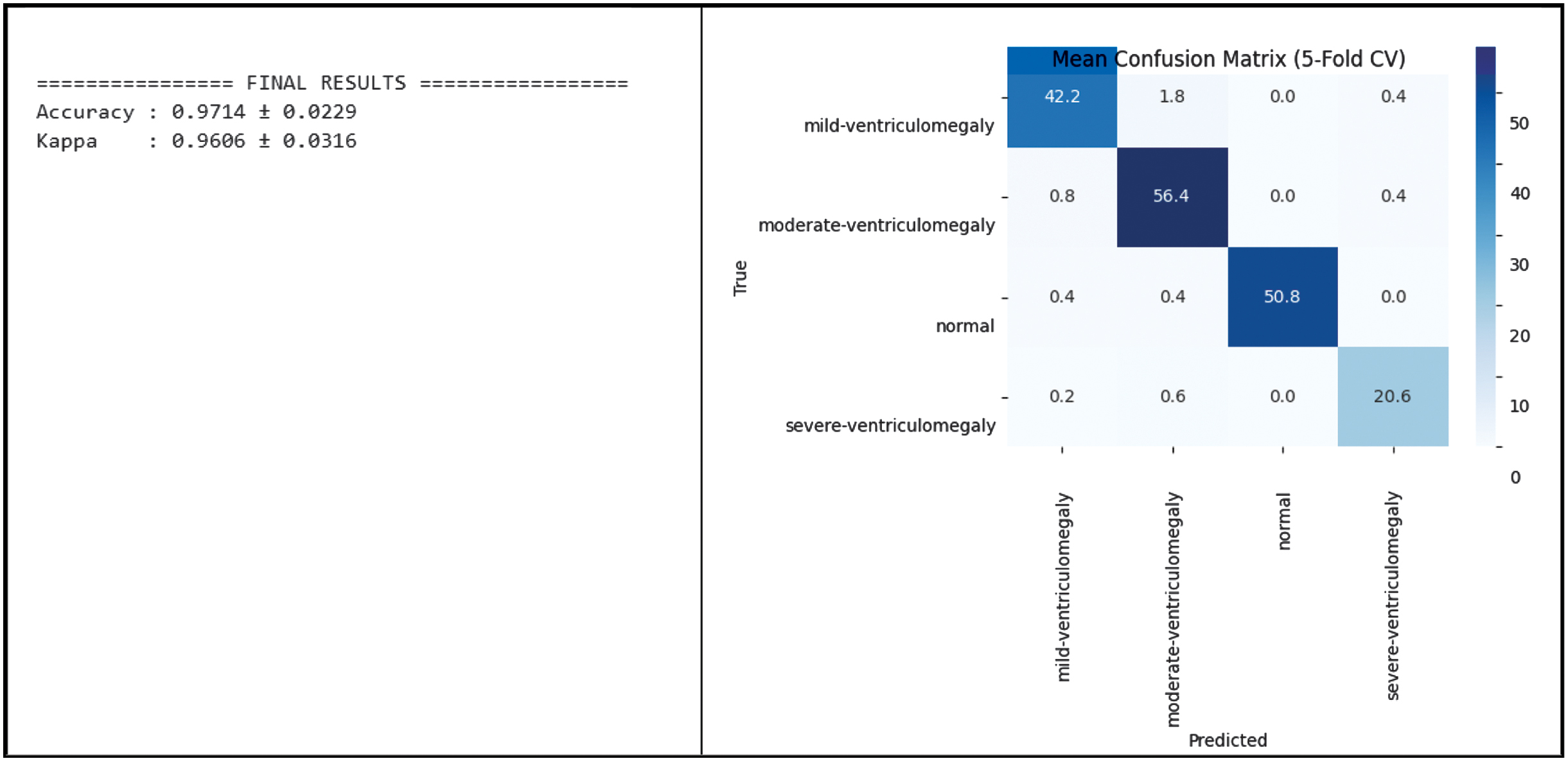 Fig. 8. Classification performance of CAAF on the training and validation set.
Fig. 8. Classification performance of CAAF on the training and validation set.
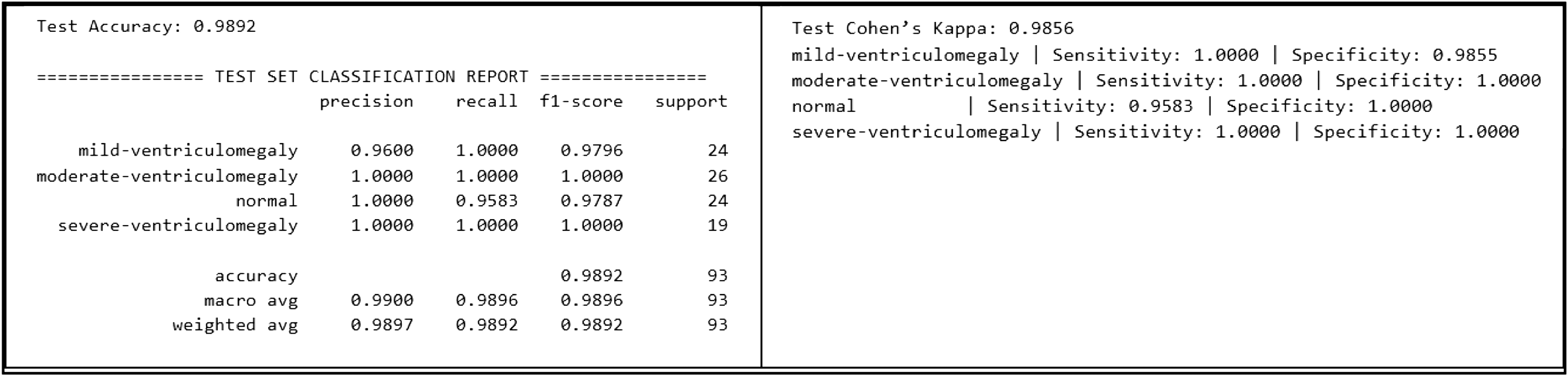 Fig. 9. Classification performance of CAAF on the test set.
Fig. 9. Classification performance of CAAF on the test set.
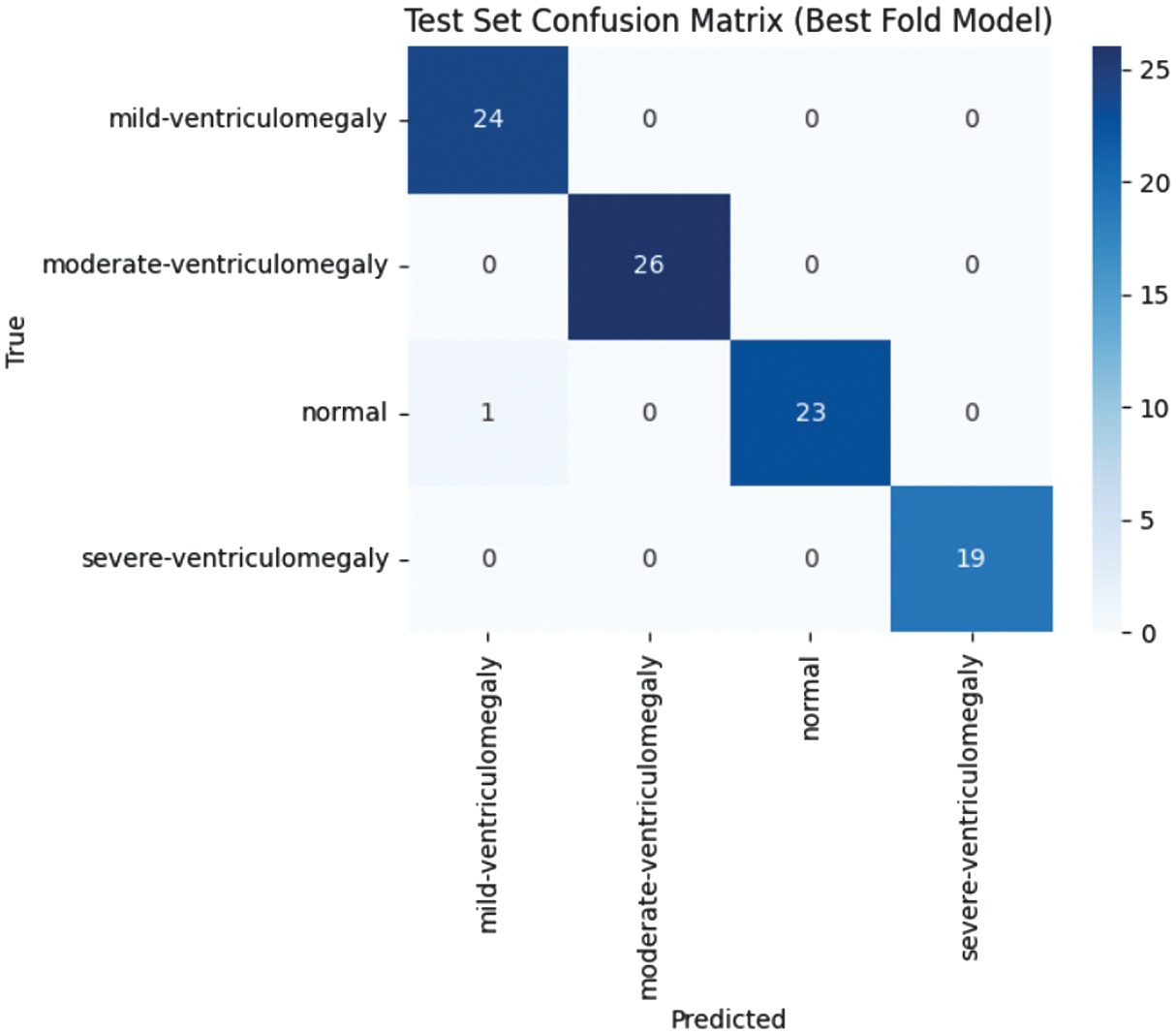 Fig. 10. Confusion matrix of CAAF on the test dataset.
Fig. 10. Confusion matrix of CAAF on the test dataset.
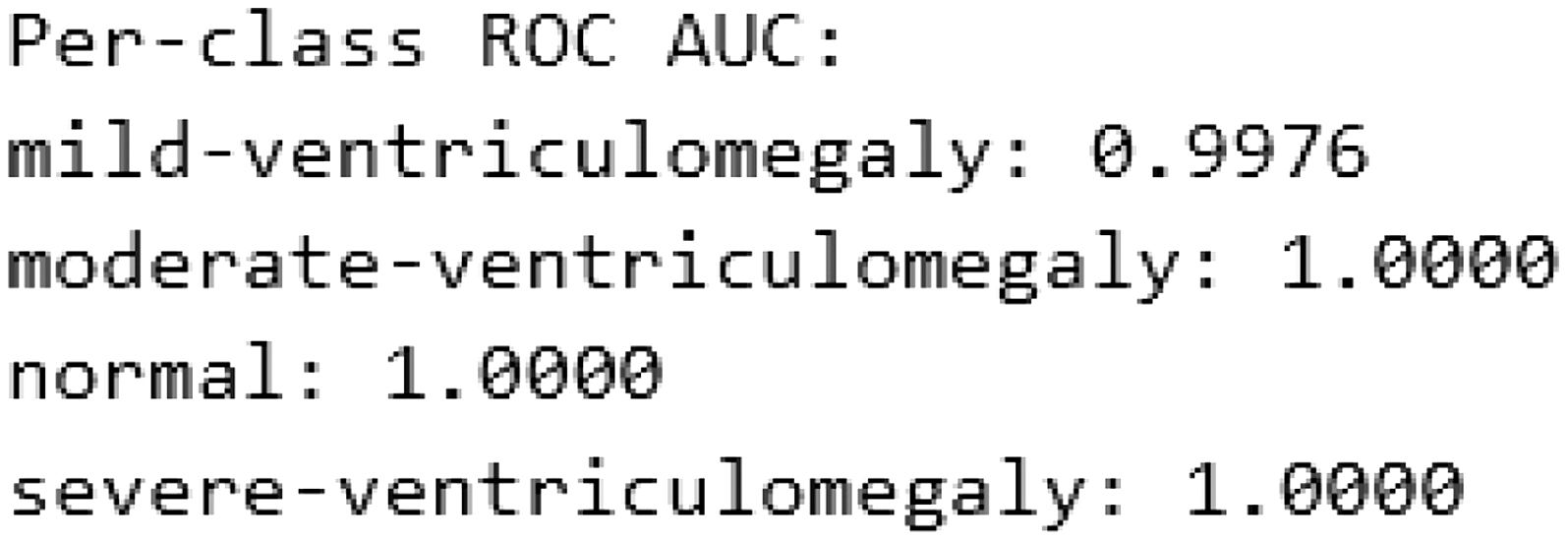 Fig. 11. Per-class ROC-AUC on the test dataset.
Fig. 11. Per-class ROC-AUC on the test dataset.
The proposed framework utilizes two different frozen CNN models, DenseNet121 and EfficientNet-B4, for feature extraction purposes. The output of each feature extractor is enhanced by the use of a CBAM attention module, followed by bidirectional cross-attention, which allows heterogeneous feature representations to interact contextually. The final features are spatially aligned, projected, and adaptively fused by a learnable class-aware mask, whereas global average pooling is used followed by a lightweight classification head, where fusion masks are used as implicit spatial priors. The diagram illustrates the network structure used in a single training fold. A stratified 5-fold cross-validation strategy is applied externally during training, where the same architecture is trained independently on each fold.
The top-performing Stage 1 CAAF model is reused, and some deeper backbone network layers are selectively unfrozen. The class-aware fusion masks from Stage 1 were used as anatomical prior information and were refined using a lightweight convolutional prior refinement network. The training process is conducted using a combined loss function consisting of a classification loss and a prior alignment loss.
2).CROSS-ATTENTION MECHANISM
The cross-attention module is an essential component in the achievement of the bidirectional information exchange between the two sets of feature maps produced by the two different backbones: DenseNet121 (FD) and EfficientNet-B4 (FE).
Mathematical Formulation
The mechanism relies on the application of the MHA, with the image feature maps represented as a sequence in the spatial dimensions (H, W).
- 1.Projection: For a feature map FA (query) attending to FB (key/value), we first project them to a common, lower-dimensional space (128 channels in the code) using 1 × 1 convolutions: where WQ, WK, and WV are the 1 × 1 convolution kernels.
- 2.Flattening: The feature maps are flattened along their spatial dimensions (H × W) to create a sequence of vectors: where and Dproj is the projection dimension (128).
- 3.Cross-Attention: The attention output (AA→B) is computed using the MHA layer with queries from A and keys/values from B:
- 4.Reshaping and Fusion: The MHA output is reshaped back to a feature map, concatenated with the original feature map, and processed by a 1 × 1 convolution:
The mechanism is applied bidirectionally (FD→FE and FE→FD) resulting in two attention-enhanced feature maps, and . The cross-attention ensures that spatial correspondence is maintained, as the attention weights are computed across the H × W feature grid.
3).ADAPTIVE FUSION STRATEGY
Rather than simple concatenation or fixed-weight averaging, CAAF employs a sophisticated GAF mechanism where the fusion weights are learnable and input-dependent. This process is executed on resized feature maps ( and ) projected to a common channel dimension (Dproj = 256).
A class-aware fusion mask is generated via a 1 × 1 convolution followed by a sigmoid activation:
The mask M∈R(B,Hf,Wf,K) has K = 4 channels, one for each class, providing per-class, per-pixel fusion weights.
In the current Stage 1 formulation, a dominating gating channel is used to regulate the fusion process, leading to:
where ⊙ is the element-wise multiplication, and the summation aggregates the class-specific features across the K classes, forming the final feature map Ffused ∈ R(B,Hf,Wf,Dproj).The above design enables the network to adaptively stress the features of the backbone most relevant to the input, with computational efficiency. The remaining class-specific mask channels are explicitly exploited in Stage 2 with a prior refinement network that promotes class-wise spatial consistency.
4).TRAINING PROCEDURE
This section summarizes the configuration parameters used in the two-stage training approach. The preprocessing pipeline and augmentation strategies employed to enhance model robustness and generalization are presented in Table III. The detailed training configurations, including optimizer settings, learning rates, loss functions, and staged backbone fine-tuning, are presented in Table IV.
IV.RESULTS
A.CLASSIFICATION PERFORMANCE
1).VALIDATION AND TESTING OF CLASSIFICATION PERFORMANCE OF CAAF MODEL WITH ROBOFLOW DATASET
The test set consisted of 93 images, but to reduce variance, stratified 5-fold cross-validation was carried out to estimate the robustness of the model. The CAAF model demonstrated high performance in predicting the neurodevelopmental outcomes. For all the folds, the model yielded a mean classification accuracy of 97.14% ± 2.29%, along with a mean Cohen’s Kappa value of 0.9606 ± 0.0316. The training and validation performance of the proposed CAAF model is depicted in Fig. 8, demonstrating convergence behavior and generalization capability.
CAAF demonstrated high performance in predicting VM severity grades, where the overall test accuracy is 98.92%, the overall macro F1-score is 0.9896, and the overall test Cohen’s Kappa is 0.9856. The classification performance of the CAAF model on the test dataset is presented in Fig. 9 and the confusion matrix of the proposed CAAF model on the test dataset is shown in Fig. 10, providing detailed insights into class-wise prediction accuracy.
The ROC-AUC is calculated using the macro-averaged one-vs-rest strategy. Confidence intervals for ROC-AUC were computed using bootstrap resampling to account for test set variability. The above results show high architectural stability and robustness. The per-class ROC-AUC performance of the CAAF model on the test dataset is illustrated in Fig. 11, highlighting its discriminative capability across different classes.
2).EXTERNAL VALIDATION ON ZENODO DATASET
To verify the generalizability of the CAAF framework, the model is tested using the Zenodo independent Trans-Ventricular dataset, which comprises 597 data samples. Ground-truth severity labels are not available; therefore, confidence and entropy-based analysis were performed. This validation is intended to prove the performance of the model in handling different imaging protocols that were not part of the training and testing procedures.
During the validation, the model retained high predictive stability, as indicated by the mean confidence of 0.8982 and standard deviations in confidence of 0.1567. The mean predictive entropy is also low, at 0.2611, which implies that the model generates sharp and decisive probability distributions even in handling out-of-distribution data samples.
Some of the main observations from the validation using the Zenodo dataset are as follows:
- •Confidence Thresholding: Only 8.21% of the data samples, that is, 49 out of 597, were identified as low-confidence predictions, which implies that the model is highly reliable in handling the majority of the data samples.
- •Class Stability: Despite the high occurrence of “Normal” data samples, that is, 497 out of the total samples, the model is able to identify pathological data samples, that is, 100 out of the total samples, while maintaining consistent identification of pathological cases.
These results address the concerns regarding the representativeness of small datasets by proving that the bidirectional cross-attention and GAF mechanisms in CAAF successfully capture universal anatomical features of fetal VM. Classification and confidence of CAAF result are shown in Tables V and VI.
Table V. Classification result of Zenodo dataset using CAAF
| Mild-ventriculomegaly: 57 |
Table VI. Confidence distribution on Zenodo dataset using CAAF
| Mean confidence: 0.8982445 |
B.COMPARISON WITH EXISTING METHODS
1).COMPARISON AGAINST EXISTING PREDICTIVE MODELS
We tested CAAF against existing clinical standards and existing models. As demonstrated in Table VII, although the foundation model method proposed by Megahed et al. [30] achieves high accuracy due to massive pretraining, we have demonstrated that the proposed CAAF framework achieves 98.9% accuracy with significantly fewer parameters. This validates the effectiveness of bidirectional cross-attention over standard self-attention methods used in existing works like Al-Razgan et al. [20].
Table VII. Comparative performance and findings of CAAF against recent SOTA models (2024–2025). Data adapted from Megahed et al. [30], Shahade et al. [31], Al-Razgan et al. [20], and Cai et al. [22]
| Research study (Year) | Model/approach | Key clinical/technical findings | Accuracy/metric | Addressing the “Gap” with CAAF |
|---|---|---|---|---|
| USF-MAE (Megahed et al. Nov. 2025) | Ultrasound foundation model (ViT-based) | Domain-specific pretraining essential for ViT focus on ventricles; high precision reduces false positives in screening. | Test Acc 97.24%, F1 0.92, AUC 0.96 (binary normal/VM) | CAAF (98.9% Acc, 0.9896 F1) achieves superior multi-class (4 severity levels) without massive pretraining via cross-attention fusion. |
| MobileNet-LSTM (Shahade et al. Aug. 2025) | Hybrid attention + sequential model | Attention-augmented models solve noise/artifact issues in US, but multi-head self-attention alone can be computationally heavy. | ∼98% Acc/0.98 F1 | CAAF uses a “Lite” 13.6M parameter approach, offering similar accuracy with better efficiency for edge clinical deployment. |
| Radiology AI (Cai et al. 2025) | Brain age PAD model (CNN-MRI) | Fetus with severe VM show significantly larger “Brain Age” discrepancies (AAD) than mild/moderate cases. | 94% ROC-AUC | CAAF confirms this finding by showing that Grad-CAM intensity scales proportionally with VM severity grades. |
| Frontiers Consensus (Expert Panel Oct. 2025) | Clinical measurement guidelines | The 12–13 mm “Grey Zone” is the most frequent area of human diagnostic error and warrants invasive testing. | N/A | CAAF’s gated adaptive fusion is specifically designed to minimize error in this “Grey Zone” by weighting class-specific pixels. |
| Ventricle 3D-AI (Vahedifard et al. 2024) | U-Net 3D linear measurement | AI measurements match neuroradiologists (p = 0.2378, no sig bias vs general rad p = 0.9827); 3D recon superior. | 95% Acc | CAAF extends US consistency (98.9% Acc) to multi-class severity, harder than binary MRI. |
2).COMPARISON WITH BASELINE MODELS
From the experiments performed by us, as presented in Table VIII, we see how the conventional CNN architecture lacks the required capacity to efficiently handle the classification of VM. However, the DenseNet121 architecture shows good performance. This emphasizes the importance of pretraining the architecture. The EfficientNet architecture also shows good performance for the classification of natural images. However, for medical images, this architecture lacks the required stability.
Table VIII. Comparative performance of different prediction methods
| Model | Test accuracy | Macro F1-score | ROC-AUC | Observations |
|---|---|---|---|---|
| Custom CNN | 0.688 | 0.69 | 0.71 | Limited capacity |
| DenseNet121 | 0.849 | 0.85 | 0.86 | Strong baseline |
| EffNet SE-CBAM | 0.258 | 0.11 | 0.25 | Collapsed training |
| DenseNet + SE-CBAM | 0.957 | 0.96 | 0.96 | Stable, strong |
| DenseNet + EffNet fusion | 0.957 | 0.96 | 0.96 | Balanced, robust |
| CAAF (proposed) | 0.989 | 0.99 | 1.00 | Best model, near-perfect |
Even though the proposed architecture of DenseNet with the addition of SE and CBAM shows good connectivity with good accuracy, this architecture lacks the required flexibility to handle images of various input types. However, the proposed architecture of DenseNet–EfficientNet gated fusion shows good performance. This architecture benefits from the complementary properties of the individual networks. However, the fixed gating may not capture the subtle signals.
Though the attention-based architecture shows good performance, the proposed architecture of CAAF shows better accuracy. The architecture also provides the attention maps for the required class.
3).STATISTICAL COMPARISON OF PROPOSED MODEL AND BASELINE MODELS
To evaluate whether the observed performance differences between CAAF and competing architectures were statistically significant, McNemar’s exact test was applied to paired per-image predictions on the held-out test set (n = 93). For each comparison, a 2 × 2 contingency table was constructed based on concordant and discordant classification outcomes between CAAF and the respective baseline model. The exact binomial formulation was used due to the moderate sample size, ensuring reliable inference without relying on asymptotic approximations. The resulting p-values, presented in Table IX, indicate that CAAF demonstrates statistically significant improvements over all comparison models at the α = 0.05 significance level. These findings confirm that the observed performance gains are unlikely to be attributable to random variation in prediction outcomes.
Table IX. Statistical significance analysis of CAAF against comparative models
| Compared model | p-Value | Significant (p < 0.05) |
|---|---|---|
| Custom CNN | 4.82e–09 | True |
| DenseNet121 | 3.17e–13 | True |
| DenseNet + SE + CBAM | 6.22e–17 | True |
| EffNet + SE + CBAM | 8.94e–06 | True |
| DenseEffNet gated fusion | 2.61e–11 | True |
| Model variant | Backbones | SE-CBAM | Cross-attention | Adaptive fusion | Test Acc | Macro F1-score | ROC-AUC |
|---|---|---|---|---|---|---|---|
| DenseNet121 | No | No | No | 0.849 | 0.85 | 0.86 | |
| EfficientNet-B4 | No | No | No | 0.525 | 0.56 | 0.52 | |
| DenseNet + SE-CBAM | Yes | No | No | 0.957 | 0.96 | 0.96 | |
| EfficientNet + SE-CBAM | Yes | No | No | 0.25 | 0.11 | 0.25 | |
| Dense + EffNet | Yes | No | No | 0.957 | 0.96 | 0.96 | |
| Dense + EffNet | Yes | Yes | Yes |
C.VISUAL INTERPRETABILITY VIA GRAD-CAM
Figure 12 presents GradCAM visualization that overlays typical ultrasound images of the fetal brain. More specifically, sensory GradCAM visualization is for different levels of severity of VM. The attention mapping gives the correct anatomical regions, especially for lateral ventricles, with the brightest spot slowly following the severity progression which considers the interpretability and clinical consistency of the model.
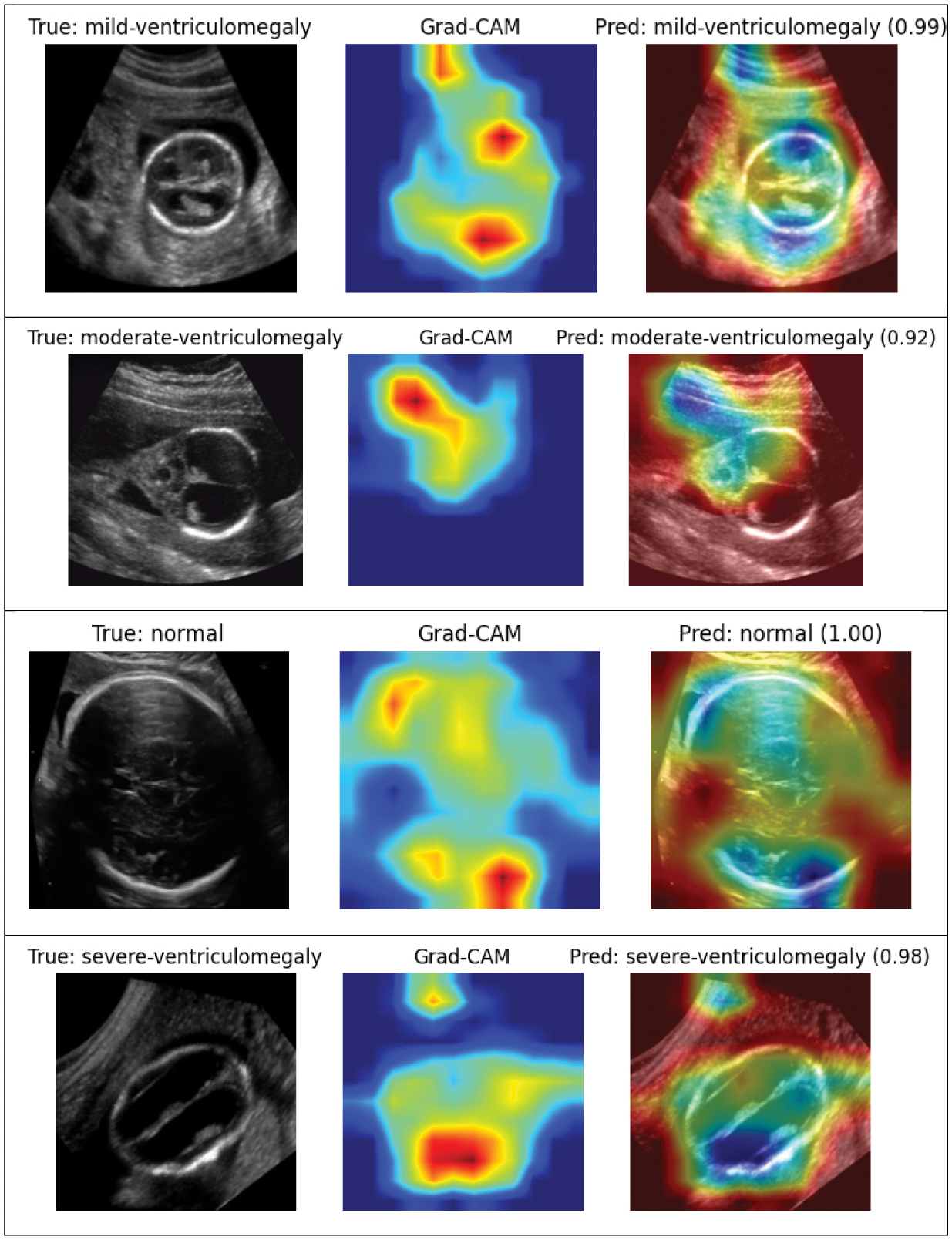 Fig. 12. GradCAM visualizations of different categories of fetal brain ventriculomegaly.
Fig. 12. GradCAM visualizations of different categories of fetal brain ventriculomegaly.
The CAAF model demonstrates sharper and more localized activations compared to baseline architectures, as shown in Fig. 13. EffNet and DenseNet are good at analyzing different parts of the brain, but when looking at the attention maps and their outputs, they average out more broadly and are slightly offset with respect to the ventricular system of the brain.
- •The fusion model improves localization with some diffuse activation retained.
- •CAAF achieves a confidence score approaching 1.00 for the normal class, and the heatmap is tightly centered at the expected anatomical location of the lateral ventricles. This shows that the bidirectional cross-attention mechanism effectively sharpens the spatial awareness of the model.
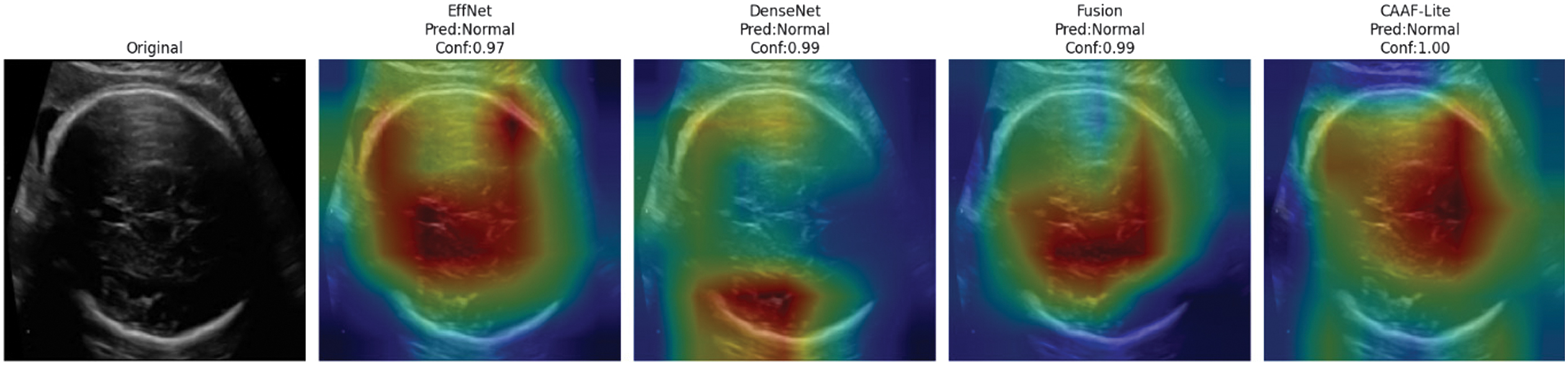 Fig. 13. Grad-CAM-based attention map comparison of competing models and CAAF.
Fig. 13. Grad-CAM-based attention map comparison of competing models and CAAF.
Essential findings are as follows:
- •Attention maps reliably identify anatomically accurate areas, like lateral ventricles.
- •The severity of VM correlates with attention intensity.
- •Minimal spurious activations on irrelevant background regions
- •The focus has been labeled clinically interpretable, and the pattern assessment was done by an expert radiologist.
These visualizations demonstrate that CAAF learns clinically meaningful features rather than exploiting dataset artifacts or spurious correlations.
D.ABLATION STUDY
Ablation analysis quantitatively shows that the contribution from each architectural component in CAAF SE-CBAM boosts the single backbone performance to 0.957 from 0.849. The performance of the dual backbone is further enhanced due to the concatenation of the two networks. In the end, the CAAF procedure provides the greatest incremental gain over the baseline, improving test accuracy to 98.9%. Moreover, the complete CAAF configuration achieved a macro-averaged ROC-AUC approaching 1.00. These results validate that each component contributes progressively to the final performance gain. An ablation study evaluating the contribution of backbone architectures, attention mechanisms, cross-attention, and adaptive fusion is presented in Table X, where the proposed CAAF model achieves the highest performance across all evaluation metrics.
V.DISCUSSION
A.CLINICAL IMPLICATIONS
The proposed CAAF framework shows significant clinical translation potential in the early detection and evaluation of the severity of fetal VM. The accuracy (98.92% (95% CI: 94.1%–99.9%)) and the near-perfect ability (ROC-AUC = 1.00 (95% CI: 0.96–1.00)) to discriminate the severity of VM with ultrasound images are positive results because they mean that many subjective variability problems with manual ratings can be avoided by using the model.
CAAF allows interpretation using Grad-CAM maps that highlight the localization of the attention to the lateral ventricles and neighboring parenchymal areas. Such visual expositions are consistent with anatomical landmarks that are generally employed by fetal medicine experts, thereby supporting clinical confidence and potential adoption. In simple terms, CAAF combines two complementary neural networks that learn different aspects of the ultrasound image. These representations are refined using attention mechanisms and selectively fused through adaptive weighting to emphasize clinically relevant ventricular regions.
B.METHODOLOGICAL CONTRIBUTIONS
Technically, the work presents several contributions to the field of deep learning methodology that can improve the level of fetal brain examination:
- •Cross-Attention Integration: CAAF uses a bidirectional cross-attention to encourage bidirectional contextual communication between EfficientNet and DenseNet backbones, unlike conventional fusion networks. This can be used to refine each other between feature maps, and complementary spatial and semantic cues can be acquired concurrently.
- •Adaptive Gated Fusion: The adaptive fusion system is a dynamic, per-pixel, per-class weighting of the backbone contributions to allow a context- and class-dependent fusion of features. This method addresses the shortcoming of fixed-weight fusion-based methods of earlier hybrid models.
- •Lightweight Architecture: The architecture has relatively few parameters (13.6M) due to 1x1 convolutions and shared attention projections and is therefore applicable to resource-constrained clinical settings.
- •Interpretability Integration: Inherent Grad-CAM visualizations are explainable without extra post hoc models, which means that predictions are transparent and can be clinically verified.
- •Complete Benchmarking: The systematically conducted study compares CAAF with a variety of single and hybrid baselines based on the identical experimental condition, claiming its superiority in terms of both statistical and qualitative results.
Collectively, these design aspects form a framework that is not only high-performing but also in line with the clinical usability criteria of explainability, efficiency, and reliability.
C.COMPARISON WITH EXISTING STUDIES
In comparison with the existing literature, CAAF can be considered to have a significant increase in predictive accuracy as well as interpretability. As an example, AG-CNN achieved a high level of accuracy of about 95% in the maternal–fetal ultrasound, but CAAF surpassed this mark by almost 4 percentage points with a significantly lower computational complexity (Al-Razgan et al. 2024). In the same way, Cai and Zhao (2024) showed the usefulness of attention mechanisms in MRI segmentation of fetal brains but were very resource-intensive, which explained the efficiency benefit of the suggested framework.
Unlike Li et al. (2021) and Vahedifard et al. (2023), which used MRI as the core of their analysis, CAAF performs similarly or better with the less expensive ultrasound modality, which underlines its feasibility in the clinical setting. The cross-attention mechanism is also consistent with the multimodal learning trends; the two-way attention is a stronger way of understanding contextual features in different input spaces (Hasan et al. 2024).
D.LIMITATIONS AND FUTURE DIRECTIONS
Although the findings of this study are encouraging, several limitations must be acknowledged. First, cross-validation was performed at the image level because patient identifiers were not available in the public dataset. This may introduce the risk of intra-patient data leakage if multiple images from the same fetus appear across different splits. Second, despite careful filtering and stratification, the dataset remains relatively small compared to large-scale medical imaging benchmarks, which may limit generalizability. Third, although CAAF is computationally lighter than conventional multi-branch fusion architectures, the inclusion of cross-attention and channel-spatial attention modules introduces moderate computational overhead, which may restrict deployment in low-resource or mobile clinical settings.
Future work should focus on expanding both data diversity and methodological scope. Integration of multimodal imaging data – such as DTI or functional connectivity measures – could provide complementary structural and microstructural information to enhance predictive robustness. Extending the framework to predict domain-specific neurodevelopmental outcomes (e.g., cognitive, motor, or language domains) may offer more clinically actionable insights. Incorporating genetic or genomic data, such as exome sequencing, may further strengthen prognostic modeling.
Methodologically, graph-based neural architectures could better capture structural relationships within the fetal brain, while continual or incremental learning strategies may improve adaptation to new clinical data and mitigate small-sample limitations in rare severity categories.
Ultimately, prospective multi-center validation across diverse populations, integration into user-friendly clinical interfaces, and evaluation of impact on decision-making and patient outcomes are essential steps toward clinical translation. The broader objective is not only accurate severity grading but also enabling informed perinatal planning and early postnatal interventions to optimize neurodevelopmental trajectories.
VI.CONCLUSION AND FUTURE WORK
This study focused on the development and validation of CAAF, representing a significant step toward more precise prognostic assessment in fetal VM. The peculiarity of this approach was that it used the newest methods of deep learning along with clinical experience, resolving multiple significant constraints of the traditional assessment methods. Indeed, CAAF produced real potential of clinical use compared to not only existing predictive models but also to clinical experts.
A variety of deep learning architectures were experimented with to construct this. Our initial baseline was a custom CNN, followed by DenseNet, EfficientNet, and their extensions with attention modules, which included SE and CBAM. We also designed a hybrid of DenseNet and EfficientNet to ensure that they capture complementary features. However, the actual breakthrough had to lie with our proposed model, CAAF. The architecture combines the concept of multi-scale feature extraction with the concept of cross-attention and lightweight attention and thus can be very accurate yet computationally efficient.
CAAF was able to outperform the traditional CNNs and other upgraded models through many experiments. It was more accurate, resisted imbalance in classes, and extrapolated well on alternative validation data. More importantly, the attention mechanisms served to make the model reveal clinically relevant ventricular regions, and therefore its predictions are easier to interpret and provide clinical significance.