I.INTRODUCTION
Diabetic foot ulcers (DFUs) represent one of the most serious and costly complications of diabetes mellitus, contributing significantly to morbidity, healthcare costs, and the risk of lower-limb amputation worldwide. Chronic wounds such as DFUs result from neuropathy, ischemia, and infection that impair the body’s natural healing process. According to global estimates, up to 25% of diabetic patients will develop a foot ulcer during their lifetime, and recurrence rates remain high even after treatment [1]. These wounds not only threaten physical mobility and quality of life but also place immense pressure on healthcare systems due to prolonged hospitalizations, frequent clinical follow-ups, and the risk of infection-related complications.
Effective management of DFUs relies on timely and accurate assessment of wound severity to determine appropriate treatment strategies. Clinicians typically evaluate ulcers using visual inspection and standardized grading systems such as the Meggitt–Wagner classification, which stratifies wounds based on depth, infection, and necrosis. However, such manual assessments are inherently subjective and prone to inter-observer variability, particularly when performed by non-specialists or under inconsistent lighting and imaging conditions [2]. The lack of objective, reproducible, and efficient diagnostic methods often results in delayed intervention, increased risk of infection, and a higher likelihood of amputation. These limitations highlight the urgent need for automated, data-driven solutions to assist clinicians in wound evaluation and monitoring.
Over the past decade, the digital transformation of healthcare has encouraged the adoption of remote diagnostic technologies, including telemedicine, to expand access to wound care. Through mobile devices or web-based platforms, patients can capture images of their wounds and receive feedback from medical professionals in real time. This approach is especially beneficial for individuals in rural or resource-limited settings where wound specialists are not readily available [3]. However, for telemedicine to be effective, it requires robust, intelligent, and interpretable AI systems capable of analyzing medical images accurately and explaining their diagnostic decisions. Artificial intelligence, particularly deep learning, offers significant promise in this domain by automating complex visual recognition tasks and supporting consistent clinical decision-making.
Deep learning models, based on convolutional neural networks (CNNs), have achieved remarkable success in diverse medical imaging tasks such as tumor detection, organ segmentation, and disease classification. In wound analysis, CNNs have been used to identify ulcer regions, classify wound types, and predict healing outcomes. Among these, segmentation models play a crucial role in isolating the wound from surrounding tissue, thereby improving the accuracy of subsequent image analysis [4]. The U-Net architecture, in particular, has become a standard framework for biomedical image segmentation due to its encoder–decoder structure and skip connections that preserve spatial information. Despite their effectiveness, segmentation models alone cannot infer clinical severity, as they focus primarily on spatial boundary delineation. Conversely, classification models, such as ResNet, DenseNet, or EfficientNet, excel in identifying image-level patterns but typically analyze full images without isolating the region of interest (ROI), introducing background noise that reduces accuracy in medical settings [5].
Existing DFU assessment models, therefore, tend to rely on single-stage deep learning architectures that perform either segmentation or classification in isolation which limits their diagnostic power. Segmentation-only approaches provide detailed wound boundaries but lack semantic interpretation, while classification-only approaches produce severity predictions without considering localized wound features [6]. Moreover, many of these models operate as black-box systems, offering high numerical performance yet providing no insight into how predictions are made. In medical contexts, where explainability and accountability are essential, this opacity hinders clinical trust and adoption. Another critical shortcoming is the neglect of ordinal relationships among severity levels in wound grading. Standard classifiers treat severity grades as independent categories, ignoring the ordered progression from mild to severe wounds. This can lead to misclassifications with major clinical implications such as labeling a Grade 4 ulcer as Grade 2 which could delay necessary medical intervention.
To address these challenges, researchers have recently begun exploring hybrid deep learning frameworks that integrate segmentation and classification into a unified pipeline. In such systems, the segmentation component isolates the lesion region, which is then passed to a classification network that determines wound severity [7]. This “image-to-mask-to-grade” paradigm enables the classifier to focus on the most relevant image areas, thereby improving accuracy and interpretability. However, even among hybrid approaches, several limitations persist. Many do not incorporate explainable AI (XAI) mechanisms, leaving clinicians unable to visualize or verify the basis of model predictions. Additionally, most hybrid architectures are designed for research settings and lack optimization for real-time telemedicine applications, where computational efficiency and low latency are vital. Despite these advances, there remains a notable absence of DFU assessment systems that simultaneously integrate ordinal-aware severity grading and XAI within a unified and clinically deployable framework.
In response to these limitations, this study proposes an explainable hybrid deep learning framework for the automated assessment of DFU severity. The system combines spatial segmentation, semantic classification, and model interpretability in a cohesive design. The segmentation module, based on the U-Net architecture, isolates the ulcer ROI from clinical images. The segmented ROI is then processed by an EfficientNet-B3 classifier, which predicts the wound severity grade using an ordinal-aware learning strategy that preserves the natural order of the Meggitt–Wagner scale [8]. To enhance transparency, Gradient-weighted Class Activation Mapping (Grad-CAM) is employed to generate heatmaps showing which regions of the image influenced the model’s decisions. These visual explanations bridge the gap between algorithmic reasoning and clinical interpretation, fostering confidence in automated assessments.
The objectives of this study are threefold: (1) to design a two-stage hybrid pipeline that effectively integrates wound segmentation and severity classification; (2) to evaluate the model’s performance on a large, annotated dataset of 6,500 DFU images using objective metrics such as Intersection over Union (IOU), Dice coefficient, accuracy, and macro-F1 score; and (3) to incorporate XAI techniques to enhance model transparency, interpretability, and suitability for clinical use.
The significance of this research lies in its contribution to the development of clinically reliable, interpretable, and computationally efficient AI systems for wound care. By bridging spatial and semantic learning, the proposed hybrid model provides a comprehensive framework for DFU analysis that addresses the shortcomings of existing single-stage models. Furthermore, its design supports real-time implementation in telemedicine and cloud-based healthcare platforms, facilitating early diagnosis, continuous wound monitoring, and equitable access to specialized care [9]. The integration of XAI ensures that both clinicians and patients can trust the system’s output, paving the way for broader acceptance of AI-assisted healthcare technologies. Ultimately, this study aims to advance the field of intelligent wound assessment by delivering a scalable and transparent deep learning solution capable of improving both clinical decision-making and patient outcomes.
The remainder of this paper is organized as follows. Section II reviews related work on deep learning-based segmentation, classification, hybrid architectures, and XAI for DFU analysis. Section III describes the dataset, proposed hybrid framework, and methodological details, including segmentation, ordinal-aware classification, and explainability components. Section IV presents experimental results and comparative evaluations, including performance analysis, benchmarking against alternative hybrid and ensemble approaches, and deployment feasibility. Section V discusses the clinical implications, limitations, and future research directions. Finally, Section I concludes the paper by summarizing the main contributions and outlining potential extensions of this work.
II.LITERATURE REVIEW
This section provides an overview of previous studies that form the conceptual foundation of this research. It expands upon the background section by examining key developments in deep learning for medical image analysis, classification of clinical images, hybrid architectures, and XAI [10]. The discussion is organized thematically and chronologically to highlight the evolution of techniques and to identify the existing research gap addressed in this study.
A.DEEP LEARNING IN MEDICAL IMAGE SEGMENTATION
Deep learning-based segmentation has become a cornerstone of medical image analysis, enabling precise delineation of pathological regions in complex clinical images. Early work introduced fully convolutional networks (FCNs) for pixel-level prediction, paving the way for automated and end-to-end segmentation frameworks [11]. A major breakthrough was the U-Net architecture, which employs an encoder–decoder structure with skip connections to preserve spatial resolution and contextual information. This design has proven particularly effective for segmenting irregular anatomical structures, including DFUs, tumors, and skin lesions.
Building on this foundation, subsequent studies proposed architectural extensions to improve segmentation accuracy and robustness. Residual U-Net (ResUNet) integrates residual connections to enhance gradient propagation and mitigate overfitting in limited medical datasets [12]. Attention U-Net further refines this approach by introducing attention gates that suppress irrelevant background regions and emphasize clinically salient features, making it well suited for complex DFU images with heterogeneous appearance. More advanced models, such as Mask R-CNN and DeepLabV3+, incorporate multi-scale feature extraction and instance-level segmentation, achieving higher IoU and Dice coefficients on challenging datasets acquired under varied lighting conditions and skin tones [13].
Despite these advances, most segmentation studies prioritize pixel-level accuracy metrics, such as IoU and Dice coefficient, without examining how segmentation quality affects downstream clinical tasks. In the context of DFUs, accurate boundary delineation alone is insufficient, as segmentation models do not encode semantic or pathological information related to wound severity. Moreover, reported performance is often based on curated datasets, limiting generalizability to real-world telemedicine images that frequently contain occlusions, illumination artifacts, and diverse wound morphologies. These limitations indicate that, while segmentation models are essential for lesion localization, they are inadequate as standalone solutions for clinically meaningful DFU assessment.
B.CLASSIFICATION NETWORKS FOR CLINICAL IMAGE ANALYSIS
Parallel to advances in segmentation, deep learning has substantially transformed the classification of medical images. Early CNNs demonstrated strong capability in identifying pathological patterns by learning hierarchical spatial and textural representations from labeled data [14]. The introduction of ResNet addressed the vanishing-gradient problem through residual connections, enabling the training of deeper and more expressive networks. Subsequent architectures, including DenseNet and EfficientNet, further improved classification performance by optimizing feature reuse and compound scaling across network depth, width, and resolution.
In the context of DFU assessment, CNN-based classifiers have been applied to predict ulcer severity, infection status, and healing potential from clinical photographs [15]. Although these models often achieve high image-level accuracy, they typically operate on full-frame images without explicitly isolating the wound ROI. As a result, classification performance can be degraded by background noise, confounding visual cues, and irrelevant anatomical features. This limitation is particularly problematic in telemedicine scenarios, where image quality and framing are highly variable.
A more critical methodological limitation is that most DFU classification approaches treat severity grading as a nominal multi-class problem, disregarding the ordinal progression inherent in clinical wound scales such as the Meggitt–Wagner system. This simplification can lead to clinically implausible errors, including large grade jumps (e.g., misclassifying advanced ulcers as mild cases), which may have significant implications for treatment planning and patient outcomes. To address this issue, ordinal regression approaches, including cumulative link models and the Consistent Rank Logits (CORAL) framework, have been proposed to explicitly model ordered relationships between severity levels [16]. These methods reduce large-grade prediction errors and improve clinical interpretability by aligning algorithmic outputs with the progressive nature of disease severity.
Despite these advances, most existing DFU classification models still suffer from two key shortcomings. First, ordinal-aware learning is not consistently integrated into state-of-the-art classification pipelines. Second, many high-performing classifiers operate as black-box systems and do not provide explainable outputs that allow clinicians to understand or validate model predictions. Consequently, despite strong performance in image-level prediction, current DFU classification models remain limited in clinical reliability, interpretability, and real-world applicability.
C.HYBRID DEEP LEARNING ARCHITECTURES
The limitations of single-stage deep learning models have motivated the development of hybrid architectures that integrate segmentation and classification within a unified pipeline [17]. Early hybrid frameworks typically adopted a two-step design in which the lesion region is first segmented and then classified according to disease severity. This “image-to-mask-to-grade” paradigm improves classification accuracy by directing the model’s attention to clinically relevant regions while reducing the influence of surrounding tissue.
Between 2019 and 2024, hybrid architectures evolved to incorporate more advanced segmentation and classification components. Studies employing U-Net or Attention U-Net for ROI extraction, followed by classifiers such as EfficientNet or ResNet, reported significant improvements in accuracy and macro-F1 scores [18]. A key advantage of these modular designs is their flexibility, allowing individual components to be optimized or updated independently without retraining the entire pipeline.
More recent work has examined error propagation within hybrid pipelines, demonstrating that segmentation quality directly affects downstream classification performance. Inaccurate or incomplete segmentation masks can lead to erroneous severity predictions, particularly for advanced ulcers occupying small or irregular regions of the image. These findings underscore the importance of accurate and stable segmentation as a prerequisite for reliable severity grading in hybrid systems.
Despite these advances, existing hybrid approaches continue to exhibit important limitations. Many treat severity grades as nominal categories rather than ordered stages of disease progression, resulting in clinically inconsistent predictions. Moreover, XAI mechanisms are often absent or incorporated only as post hoc visualization tools, limiting clinicians’ ability to understand and validate model decisions [19]. Consequently, while hybrid segmentation–classification pipelines improve grading accuracy by focusing on lesion regions, most existing approaches fail to integrate ordinal-aware learning with explainable mechanisms, leaving a critical gap in transparent and clinically consistent DFU assessment suitable for real-world clinical decision support.
D.XAI AND INTEGRATION WITH TELEMEDICINE
As deep learning models become increasingly complex, the demand for XAI in healthcare has grown substantially. Explainability enhances transparency in model decision-making, enabling clinicians to verify whether algorithmic attention aligns with medically relevant features and established clinical reasoning [20]. Among the most widely adopted visualization techniques, Grad-CAM projects class-specific gradients onto convolutional feature maps to generate heatmaps that highlight image regions most influential in a given prediction. Grad-CAM and its extensions have been successfully applied across various medical imaging tasks to improve interpretability and clinician confidence.
Subsequent methods, such as Score-CAM, refine gradient-based approaches by weighting activation maps using class scores, resulting in more stable and visually coherent explanations. Beyond visualization, XAI techniques have been incorporated into clinical auditing, model validation, and human-in-the-loop decision-making frameworks, reinforcing accountability and trust in automated systems. The growing body of evidence indicates that transparent and interpretable models are essential not only for regulatory compliance but also for facilitating clinical acceptance of AI-assisted diagnostics.
In parallel, advances in telemedicine, cloud computing, and the Internet of Things (IoT) have transformed healthcare delivery by enabling remote monitoring and real-time decision support. Edge- and cloud-based AI systems can now perform near real-time image analysis, supporting wound monitoring, triage, and treatment guidance in geographically dispersed or resource-limited settings [21]. Deep learning pipelines optimized for low-latency inference can be integrated into smartphone or web-based platforms, allowing clinicians to assess patient-submitted images without requiring in-person visits.
Despite these technological advances, significant limitations remain. In telemedicine environments, where clinicians must rely on remote assessments and automated recommendations, AI systems must balance efficiency with transparency. Black-box models that deliver rapid but opaque predictions pose ethical, clinical, and medico-legal risks. Moreover, explainability is often evaluated in isolation and rarely integrated with task-specific constraints such as ordinal severity modeling, which is critical for clinically consistent disease grading. Current systems frequently optimize inference speed or accuracy independently, rather than jointly addressing transparency, reliability, and scalability.
These limitations underscore the need for unified hybrid frameworks that integrate XAI with ordinal-aware severity modeling and efficient deployment strategies. Such frameworks are essential for supporting trustworthy, real-time DFU assessment in telemedicine settings and for aligning AI-assisted diagnostics with clinical workflows and decision-making requirements.
E.SYNTHESIS AND RESEARCH GAP
The reviewed literature demonstrates substantial progress in deep learning-based medical image analysis over the past decade. Notable advancements include improved segmentation accuracy through attention and multi-scale architectures, enhanced classification performance enabled by efficient and ordinal-aware networks, and increasing emphasis on explainability to facilitate clinical adoption [22]. Despite these developments, several critical challenges remain insufficiently addressed in the context of DFU assessment.
First, segmentation and classification are often treated as independent stages, limiting end-to-end interpretability and preventing systematic analysis of how localization quality influences severity grading. Second, many DFU classification systems continue to model severity levels as nominal categories rather than ordered stages of disease progression, leading to clinically inconsistent predictions. Third, although XAI techniques such as Grad-CAM have been proposed, their integration into DFU severity assessment remains limited, and visual explanations are rarely aligned with or validated against clinical reasoning. Finally, relatively few studies assess model latency, scalability, and robustness in real-world telemedicine settings, where efficiency, transparency, and reliability are equally essential.
Addressing these limitations requires a unified approach that moves beyond isolated improvements in individual components. In particular, the absence of frameworks that jointly integrate precise lesion segmentation, ordinal-aware severity grading, and explainable decision-making represents a key gap in current DFU assessment research.
This study addresses this gap by proposing an explainable hybrid deep learning framework that integrates U-Net-based segmentation, ordinal-aware EfficientNet-B3 classification, and Grad-CAM visualization within a single pipeline [23]. The proposed model is explicitly designed to achieve high accuracy, interpretability, and computational efficiency, enabling trustworthy and real-time deployment in telemedicine and cloud-based healthcare systems.
F.COMPARISON WITH ALTERNATIVE HYBRID DEEP LEARNING FRAMEWORKS
Several hybrid deep learning frameworks have been proposed for DFU analysis, typically combining a segmentation backbone with a classification network to improve grading accuracy. Common configurations include U-Net paired with ResNet, DenseNet, or VGG-based classifiers, as well as attention-enhanced segmentation combined with conventional categorical classifiers. While these approaches report improvements over single-stage models, their comparative advantages and limitations warrant critical examination.
Compared with U-Net + ResNet hybrids, which emphasize deep residual feature extraction, the proposed U-Net + EfficientNet-B3 framework achieves comparable or superior classification accuracy with reduced computational complexity due to EfficientNet’s compound scaling strategy. DenseNet-based hybrids benefit from feature reuse but often incur higher memory overhead, limiting deployment feasibility in telemedicine and mobile environments. VGG-based hybrids, although structurally simple, typically underperform in both accuracy and efficiency due to their large parameter count and lack of modern optimization mechanisms.
A key distinction of the proposed framework lies in its integration of ordinal-aware learning, which is absent from most existing hybrid DFU systems. Prior hybrid models generally treat severity grading as a nominal classification task, leading to clinically inconsistent errors between non-adjacent grades. By contrast, the use of consistent rank logits in the EfficientNet-B3 classifier enforces ordinal constraints aligned with the Meggitt–Wagner scale, reducing large-grade misclassifications that are common in alternative hybrids.
Furthermore, while some hybrid frameworks incorporate post hoc visualization, explainability is rarely treated as a core design objective. In contrast to existing hybrids that prioritize numerical accuracy alone, the proposed pipeline embeds Grad-CAM-based visual explanations directly into the classification stage, enabling verification of model attention and supporting clinician trust. This combination of segmentation-guided input, ordinal-aware grading, and explainable output differentiates the proposed approach from previously reported hybrid DFU frameworks.
Overall, the comparative analysis indicates that the U-Net + EfficientNet-B3 configuration offers a more balanced trade-off between accuracy, interpretability, and computational efficiency than alternative hybrid architectures, making it better suited for real-world telemedicine deployment.
III.METHODS
This section describes the dataset, architecture design, training strategy, and evaluation procedures used in developing the proposed hybrid deep learning framework for DFU assessment. All methods are presented in sufficient detail to enable replication.
A.DATASET
A total of 6,500 high-resolution DFU images were collected from several hospitals and outpatient centers under institutional ethical approval. Each image depicts plantar or dorsal views of diabetic feet under varying illumination, skin pigmentation, and ulcer morphology, ensuring a diverse representation of real-world conditions.
The dataset used in this study consists of 6,500 clinical images of DFUs collected from multiple hospitals and outpatient wound-care centers through institutional collaboration. All data were obtained under approved institutional review procedures, with patient identifiers removed prior to analysis to ensure anonymity and compliance with ethical standards for secondary clinical data use. The study did not involve direct patient interaction, and informed consent was waived in accordance with institutional and national research ethics guidelines. Due to patient privacy and data-sharing restrictions imposed by participating institutions, the dataset is not publicly available; however, it may be accessed for academic research purposes upon reasonable request and subject to institutional approval.
Each sample includes two forms of annotation:
- •Segmentation masks: binary images labeling ulcer pixels as 1 and background pixels as 0. Masks were manually drawn by trained medical annotators and verified by two wound-care specialists. Manual annotation remains the clinical gold standard for medical image segmentation.
- •Severity grades: ordinal labels from Grade 0 to 5 following the Meggitt–Wagner classification system, assigned by at least two certified clinicians. Discrepancies were resolved by consensus review to ensure consistency.
The dataset is divided using stratified sampling to preserve class balance: 70% for training, 15% for validation, and 15% for independent testing. To enhance generalization, on-the-fly data augmentation is performed only on the training set, including random horizontal/vertical flips, ±20° rotations, brightness/contrast jitter, and Gaussian noise addition. The original class distribution was maintained to avoid bias.
All experiments are conducted in Python 3.10 using TensorFlow 2.9 and Keras 2.9 on a workstation equipped with an NVIDIA RTX 3060 GPU (12 GB VRAM), Intel Core i7 CPU, and 32 GB RAM. To support reproducibility, the implementation details, training configurations, and model weights will be made available to the research community upon reasonable request, subject to institutional and ethical approval, with a public code repository planned for release following manuscript acceptance.
B.HYBRID DEEP LEARNING PIPELINE
The proposed framework (Fig. 1) integrates segmentation, classification, and interpretability into a unified deep learning pipeline for automated DFU severity assessment. The system follows four sequential stages:
- 1.Image input,
- 2.Wound segmentation via U-Net,
- 3.ROI extraction and severity classification using EfficientNet-B3 enhanced by Grad-CAM visualization.
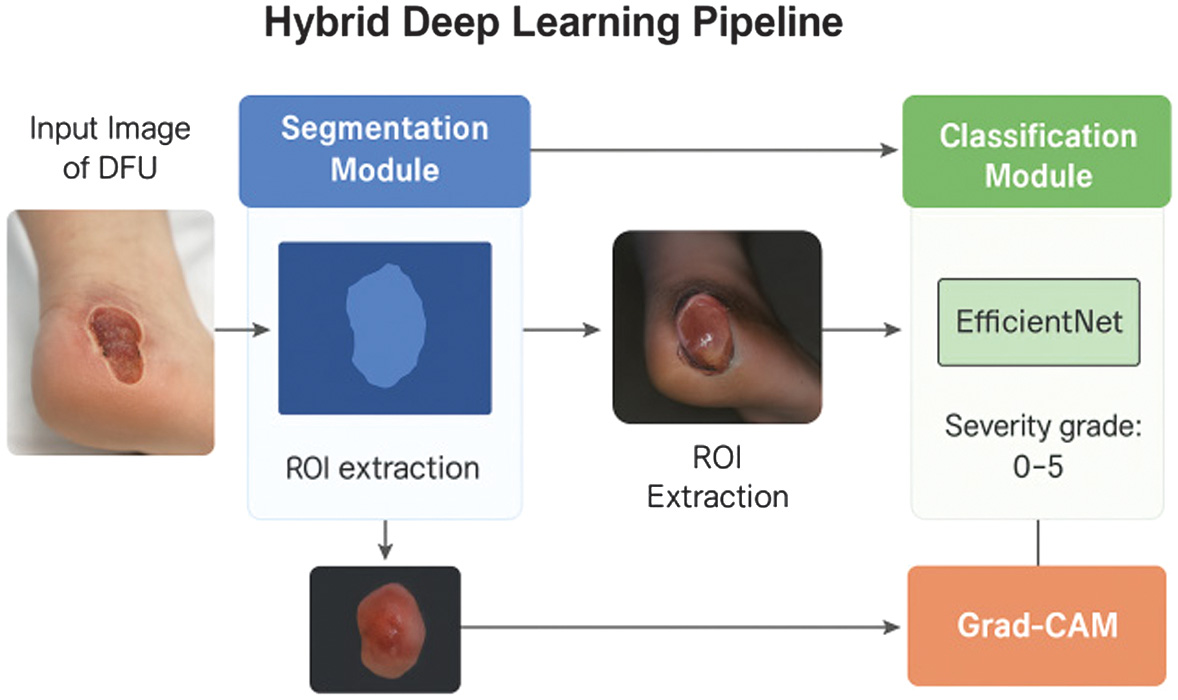 Fig. 1. Overview of the proposed hybrid deep learning framework for diabetic foot ulcer (DFU) assessment.
Fig. 1. Overview of the proposed hybrid deep learning framework for diabetic foot ulcer (DFU) assessment.
Each component was designed to preserve spatial accuracy, semantic richness, and transparency while maintaining computational efficiency.
The framework implements a four-stage pipeline: (1) acquisition of a raw clinical foot image; (2) automated ulcer segmentation using a U-Net-based model to generate a binary wound mask; (3) ROI extraction guided by the segmentation output; and (4) severity classification using an EfficientNet-B3 network with ordinal-aware learning. Grad-CAM visualization is applied to highlight image regions that most influence the predicted wound grade (0–5), enabling visual verification of model reasoning and supporting transparent, clinically meaningful DFU assessment.
1).SEGMENTATION MODULE
The first stage automatically isolates the ulcer region from surrounding tissue, which is essential for suppressing background interference and improving downstream classification accuracy. As shown in Fig. 2, the segmentation network successfully separates the lesion area across wounds of varying color, size, and illumination.
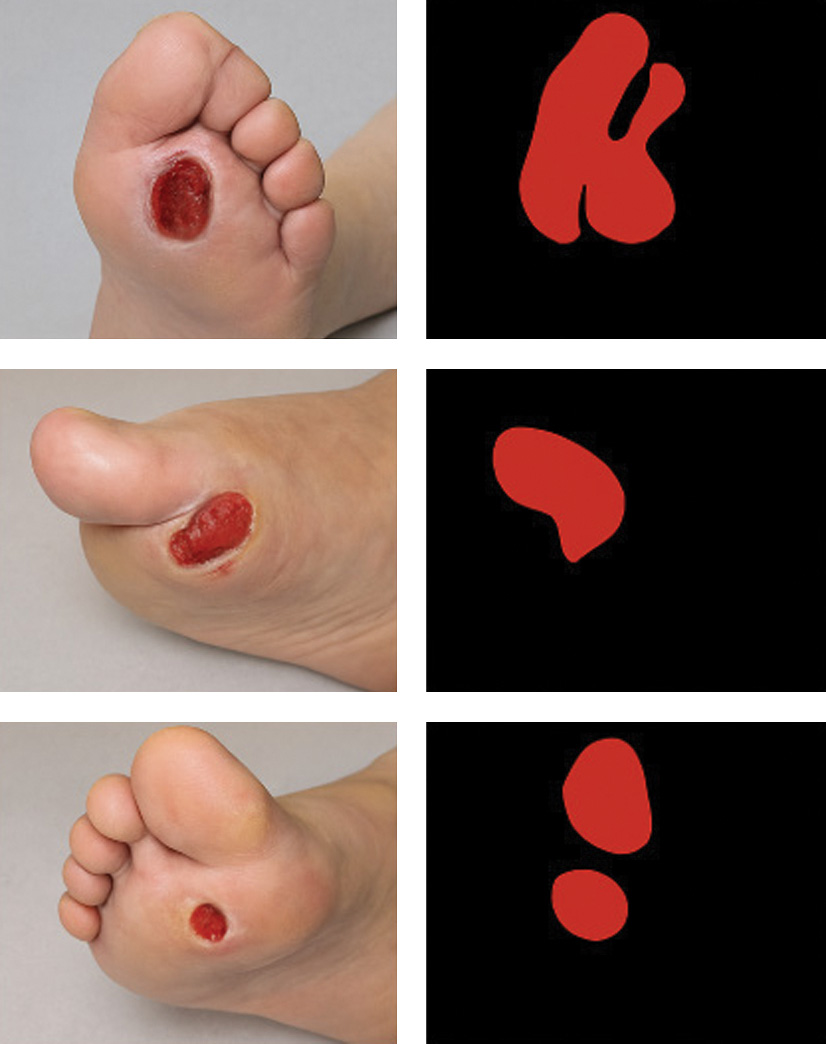 Fig. 2. Sample DFU images with ground-truth masks.
Fig. 2. Sample DFU images with ground-truth masks.
The U-Net architecture was selected for ulcer segmentation due to its strong suitability for medical imaging tasks involving limited and heterogeneous datasets. Its encoder–decoder structure with skip connections enables precise localization of lesion boundaries while preserving fine-grained spatial information, which is critical for accurately delineating DFUs with irregular shapes and variable appearance. Compared with more complex architectures such as DeepLabV3 + or Mask R-CNN, U-Net offers a favorable balance between boundary precision and computational efficiency, making it well suited for real-time telemedicine applications. Additionally, U-Net has demonstrated robust performance across a wide range of biomedical segmentation benchmarks, particularly in scenarios with moderate dataset sizes, where deeper or multi-branch models may be prone to overfitting or increased inference latency. These characteristics make U-Net an appropriate and reliable choice for the segmentation stage of the proposed hybrid framework.
A U-Net architecture was selected as the baseline because of its proven reliability in biomedical image segmentation. It employs an encoder–decoder structure in which high-resolution feature maps from early layers are concatenated with deeper semantic features through skip connections. This fusion allows the model to capture both global context and fine local boundaries. To further refine spatial focus, an Attention U-Net variant was implemented; attention gates within its skip connections learn to suppress non-informative background pixels and emphasize clinically relevant regions.
All images were resized to 256 × 256 pixels and normalized to the [0, 1] range. During training, real-time augmentation (random flips, ±20° rotations, brightness, and contrast adjustments) was applied to increase robustness. The segmentation model was trained using a composite loss function combining Dice Loss and Binary Cross-Entropy (BCE):
Dice Loss enhances sensitivity to small or irregular lesions, while BCE ensures pixel-wise accuracy; together they counter class imbalance. Optimization used the AdamW optimizer (learning rate =1 × 10−4, batch size = 8, 100 epochs). Segmentation quality was measured by IoU and the Dice coefficient.
Each row displays the original DFU image, corresponding binary mask, and the overlay of the mask on the original image. The results demonstrate the model’s ability to accurately delineate ulcer boundaries despite lighting variation and complex skin textures.
2).CLASSIFICATION MODULE
Following segmentation, the cropped ROI is sent to the classification network. Removing unrelated background skin allows the classifier to concentrate on wound morphology, texture, and color key features for determining clinical severity.
The classifier uses EfficientNet-B3, a CNN that applies compound scaling across network depth, width, and resolution to achieve an optimal balance between accuracy and efficiency. The network was initialized with ImageNet weights and fine-tuned on the DFU dataset to leverage transfer learning. Prior to training, ROIs were resized to 224 × 224 pixels, normalized, and enhanced using Contrast-Limited Adaptive Histogram Equalization (CLAHE) to improve visualization of necrotic and granulated tissue.
To incorporate the ordered nature of the Meggitt–Wagner grading scale, training employed the CORAL ordinal-regression loss. This cumulative-link approach models ordered thresholds between severity levels, ensuring that predictions respect clinical progression (Grade 3 is closer to Grade 2 than Grade 5). The loss penalizes large-grade errors more heavily than minor deviations, thereby improving both numerical accuracy and clinical interpretability.
The final output includes the predicted severity grade (0–5) and a Grad-CAM-based heatmap highlighting the discriminative regions influencing the prediction. As shown in Fig. 3, the heatmaps generated by the classifier concentrate on the ulcer core and border areas that align with clinician focus demonstrating the model’s interpretability and reliability.
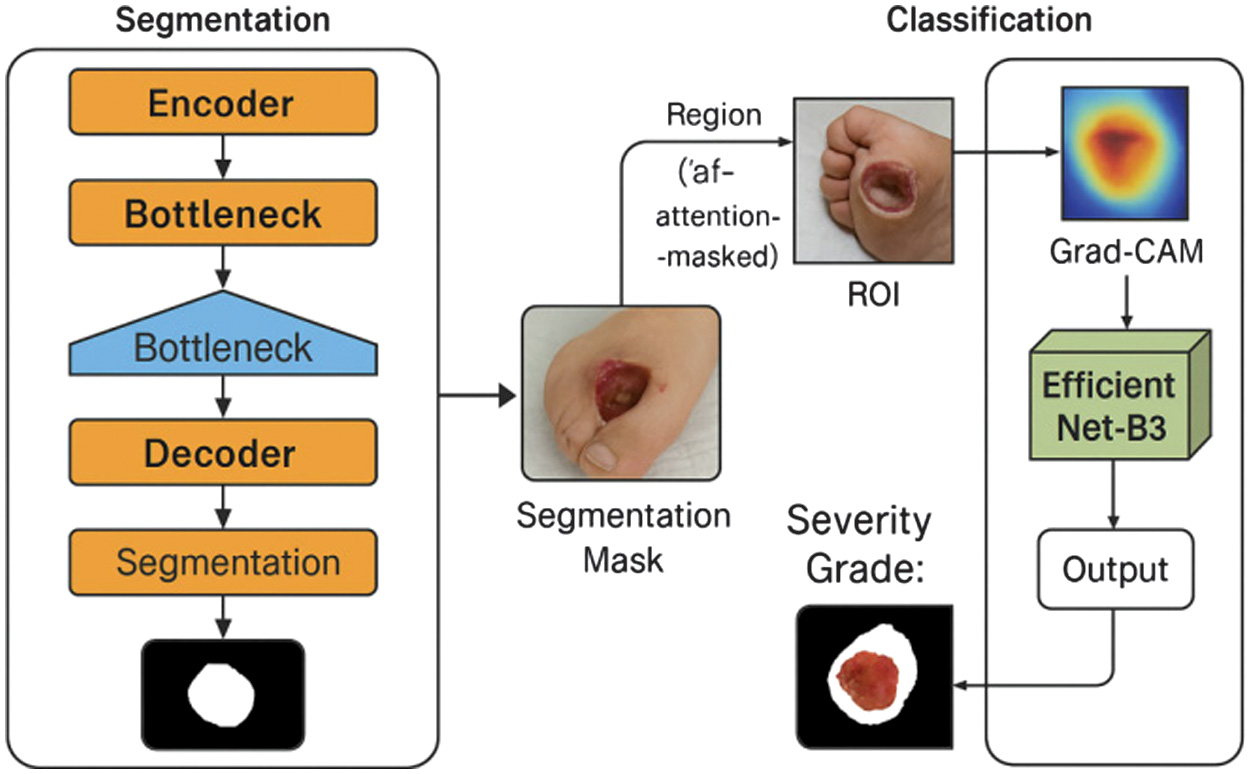 Fig. 3. Hybrid pipeline workflow and Grad-CAM visualization.
Fig. 3. Hybrid pipeline workflow and Grad-CAM visualization.
Figure 3 illustrates each processing stage: (a) input image, (b) segmentation mask, (c) ROI extraction, (d) classification output, and (e) Grad-CAM overlay. Warmer colors represent higher model attention over the ulcer site.
3).EXPLAINABILITY MODULE
Model interpretability is achieved using Grad-CAM. Grad-CAM projects the gradients of the target class onto the final convolutional feature maps, producing a coarse localization map that highlights discriminative regions. Warmer colors in the heatmap correspond to higher model attention.
For each test image, the generated Grad-CAM overlay is visually inspected to confirm whether the network focused on clinically meaningful ulcer regions (edges, necrotic centers, erythematous zones). This qualitative analysis enhances transparency and supports human-in-the-loop validation during telemedicine deployment.
C.TRAINING AND OPTIMIZATION STRATEGY
Both modules are trained independently and then integrated for end-to-end evaluation. All models use the AdamW optimizer with default β-parameters (β1 = 0.9, β2 = 0.999) and decoupled weight decay = 1 × 10−4. The learning rate is reduced by a factor of 0.1 after 10 epochs of plateau. Early stopping was applied when validation loss did not improve for 10 consecutive epochs. Data augmentation strategy:
- •Segmentation: random rotations (±20°), flips, and color jitter.
- •Classification: random brightness/contrast shifts and CLAHE normalization.
These augmentations simulate real clinical variability (lighting, orientation, and camera noise) and improve robustness. Training duration: approximately 6 hours for segmentation and 5 hours for classification on the specified hardware. Model checkpoints are stored after each epoch for reproducibility.
D.EVALUATION METRICS
To ensure objective and comprehensive performance assessment, four groups of metrics were applied:
- 1.Segmentation performance–evaluated by
- ○IOU: Dice coefficient: where and represent predicted and ground-truth masks, respectively. High scores indicate strong spatial overlap and boundary fidelity.
- 2.Classification performance–evaluated by
- ○Accuracy, precision, recall, and macro-F1 score, which balance per-class performance on imbalanced medical datasets;
- ○Confusion matrix to visualize inter-grade misclassifications and verify ordinal consistency.
- 3.Hybrid pipeline evaluation–classification accuracy is compared under two conditions: (a) with segmentation-based ROI input and (b) with unsegmented full images. The comparison quantifies the contribution of lesion localization to grading accuracy. Additionally, error propagation analysis examines how segmentation quality affects classification results, following methods outlined in prior hybrid studies.
- 4.Explainability evaluation qualitative analysis of Grad-CAM heatmaps assessed whether model attention corresponded to clinically relevant wound regions. Heatmaps showing focus on ulcer centers and edges were considered valid explanations, whereas diffuse or irrelevant activations indicated potential model bias.
All quantitative metrics were averaged over the test set with 95% confidence intervals computed via bootstrapping.
E.IMPLEMENTATION AND REPRODUCIBILITY
All experiments are executed in a controlled environment using the following configurations:
- •Programming language: Python 3.10
- •Libraries: TensorFlow 2.9, Keras 2.9, OpenCV 4.7, NumPy 1.26, Matplotlib 3.8
- •Hardware: NVIDIA RTX 3060 GPU (12 GB), Intel Core i7 CPU, 32 GB RAM
- •Operating system: Windows 11 64-bit
Random seeds are fixed across TensorFlow, NumPy, and Python to ensure deterministic outcomes. All training and evaluation scripts, along with preprocessing pipelines and model weights, can be provided by the authors for academic reproduction. No external code packages beyond the versions listed were used. Pre-registration was not required, as this work does not involve human or animal experimentation but utilizes anonymized secondary clinical images with institutional clearance.
IV.RESULTS AND DISCUSSION
This section presents and interprets the results obtained from the hybrid deep learning framework for DFU assessment. Quantitative metrics, visual examples, and comparative analyses are used to demonstrate performance improvements, clinical relevance, and interpretability [24]. The discussion also positions these findings within the context of previous studies and highlights the implications for real-world telemedicine applications.
A.SEGMENTATION PERFORMANCE
The segmentation module forms the basis of the hybrid architecture, enabling lesion-focused classification. Its precision determines how effectively the classifier isolates meaningful features from noisy image backgrounds.
Using IOU and Dice coefficient metrics, the baseline U-Net achieved an IOU of 0.843 and a Dice coefficient of 0.912, demonstrating strong alignment between predicted masks and expert annotations [25]. When the Attention U-Net variant was applied, these values increased to 0.861 (IOU) and 0.924 (Dice), showing measurable gains in boundary precision and robustness against texture variations.
These metrics are comparable with high-performing biomedical segmentation networks in other medical domains such as skin lesion detection (Dice ≈0.92) and diabetic retinopathy segmentation (Dice≈0.90) indicating the adaptability of U-Net-based architectures to diabetic foot imaging. The improvement from attention mechanisms can be attributed to the model’s ability to assign higher weights to informative pixels while suppressing irrelevant context, as illustrated in Fig. 4.
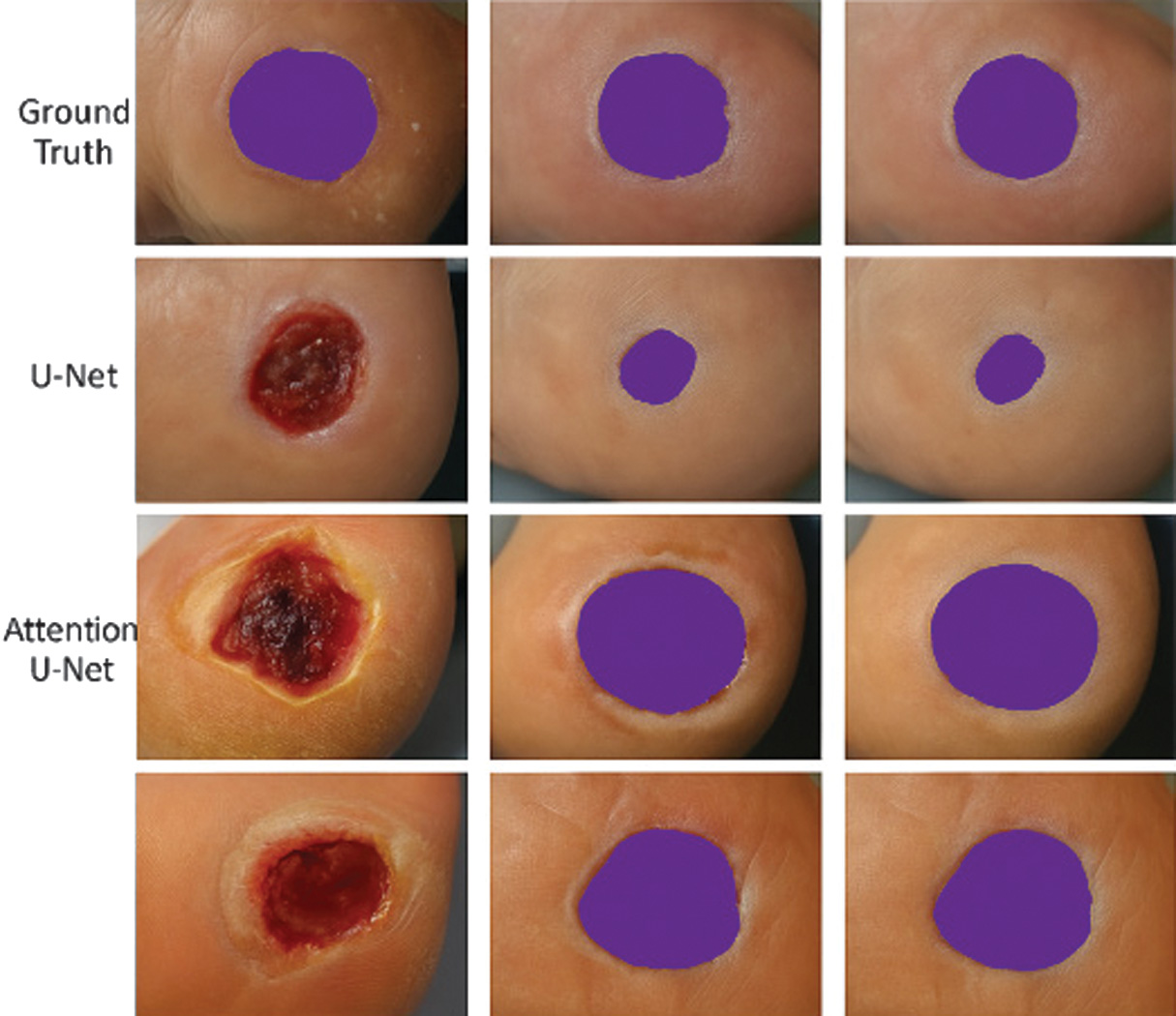 Fig. 4. Segmentation results: Ground truth vs. predicted masks.
Fig. 4. Segmentation results: Ground truth vs. predicted masks.
Visual comparison between real DFU samples and corresponding predictions from U-Net and Attention U-Net. Each column displays a distinct ulcer case, showing how the attention mechanism refines wound boundary detection under challenging conditions such as lighting variation, overlapping toes, or partially occluded lesions.
Qualitative analysis shows that segmentation accuracy is most affected by ulcer appearance variability; bright reflections from moist tissue or darkened necrotic regions can confuse the model [26]. Nonetheless, the Attention U-Net demonstrated stable performance even under such circumstances, producing continuous and anatomically consistent contours.
This high segmentation accuracy ensures that the subsequent classification network receives accurate ROI inputs, which improves both computational efficiency and clinical interpretability. Accurate wound boundary localization also facilitates longitudinal monitoring, allowing automatic computation of wound area, perimeter, and healing rate over time metrics that are essential in diabetic care.
B.CLASSIFICATION PERFORMANCE
The classification module predicts DFU severity grades (0–5) according to the Meggitt–Wagner scale. The model achieved an overall accuracy of 92.3% and a macro-F1 score of 0.904, demonstrating high predictive performance across all severity levels.
When compared with baseline architectures, EfficientNet-B3 outperformed ResNet-50 (accuracy = 89.2%, F1 = 0.872) and VGG16 (accuracy = 86.7%, F1 = 0.841) [27]. These results confirm the effectiveness of compound scaling in EfficientNet, which optimizes network depth, width, and resolution simultaneously to improve feature extraction without excessive computational cost.
Performance gains are most pronounced in mid- to late-stage ulcers (Grades 3–5), where morphological heterogeneity is highest. These cases often involve complex textural cues such as necrotic patches, deep fissures, or infection indicators that require robust multi-scale feature extraction. The classifier also performed well for lower grades, distinguishing mild abrasions (Grade 1) from healthy tissue (Grade 0) with minimal false positives in Fig. 5.
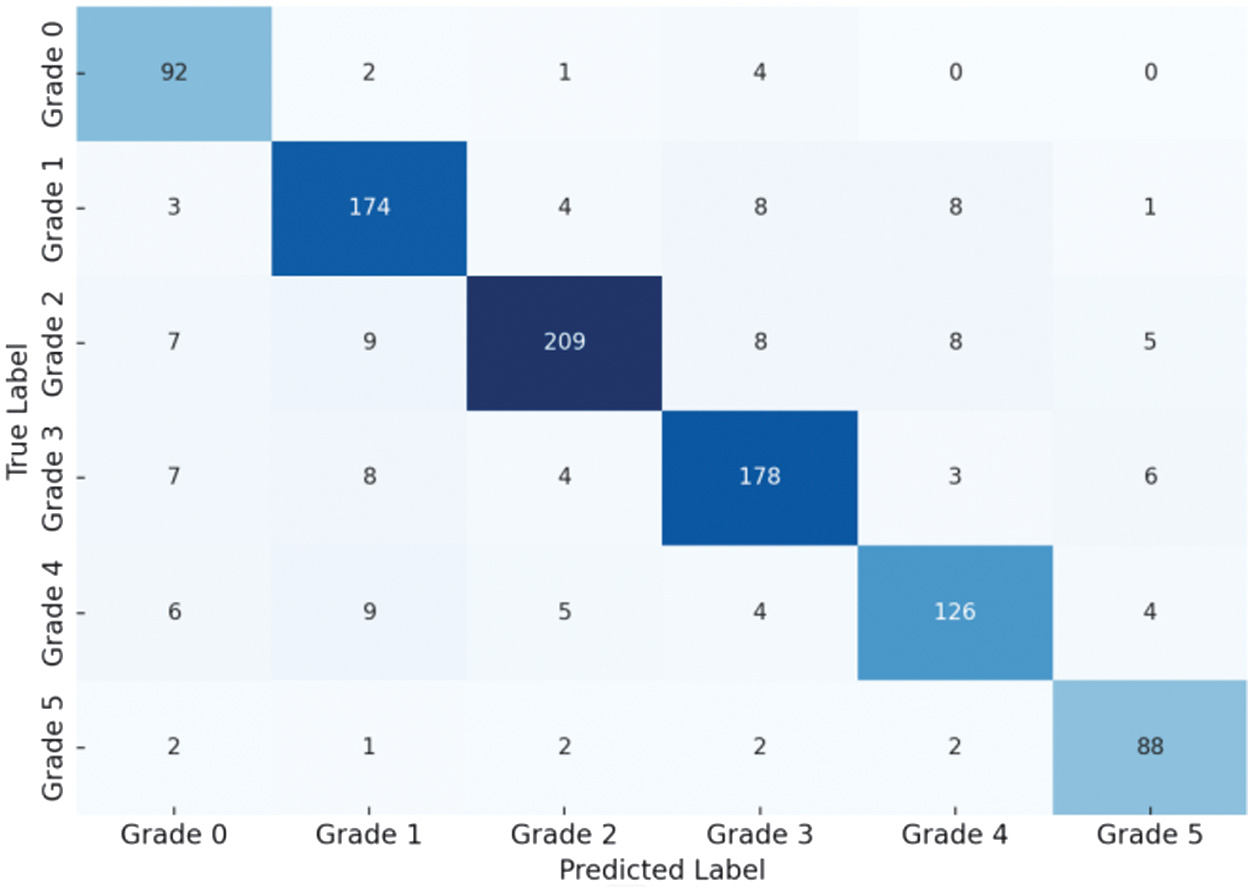 Fig. 5. Confusion matrix illustrating the performance of the ordinal-aware EfficientNet-B3 classifier for diabetic foot ulcer (DFU) severity grading.
Fig. 5. Confusion matrix illustrating the performance of the ordinal-aware EfficientNet-B3 classifier for diabetic foot ulcer (DFU) severity grading.
Rows represent the ground-truth severity grades (0–5), while columns indicate the predicted grades. The strong concentration of values along the diagonal indicates high classification accuracy, whereas the majority of misclassifications occur between adjacent grades (e.g., 2 ↔ 3 and 3 ↔ 4), demonstrating ordinal consistency and validating the effectiveness of the CORAL loss.
The ordinal-aware training proved crucial. Standard categorical cross-entropy loss treats severity classes as independent categories, allowing large jumps between unrelated grades. By contrast, CORAL loss enforces hierarchical consistency, aligning model behavior with the progressive nature of diabetic wound pathology [28]. This approach penalizes distant misclassifications more heavily, thus reducing clinically unacceptable grade reversals (1 → 5) in Fig. 6.
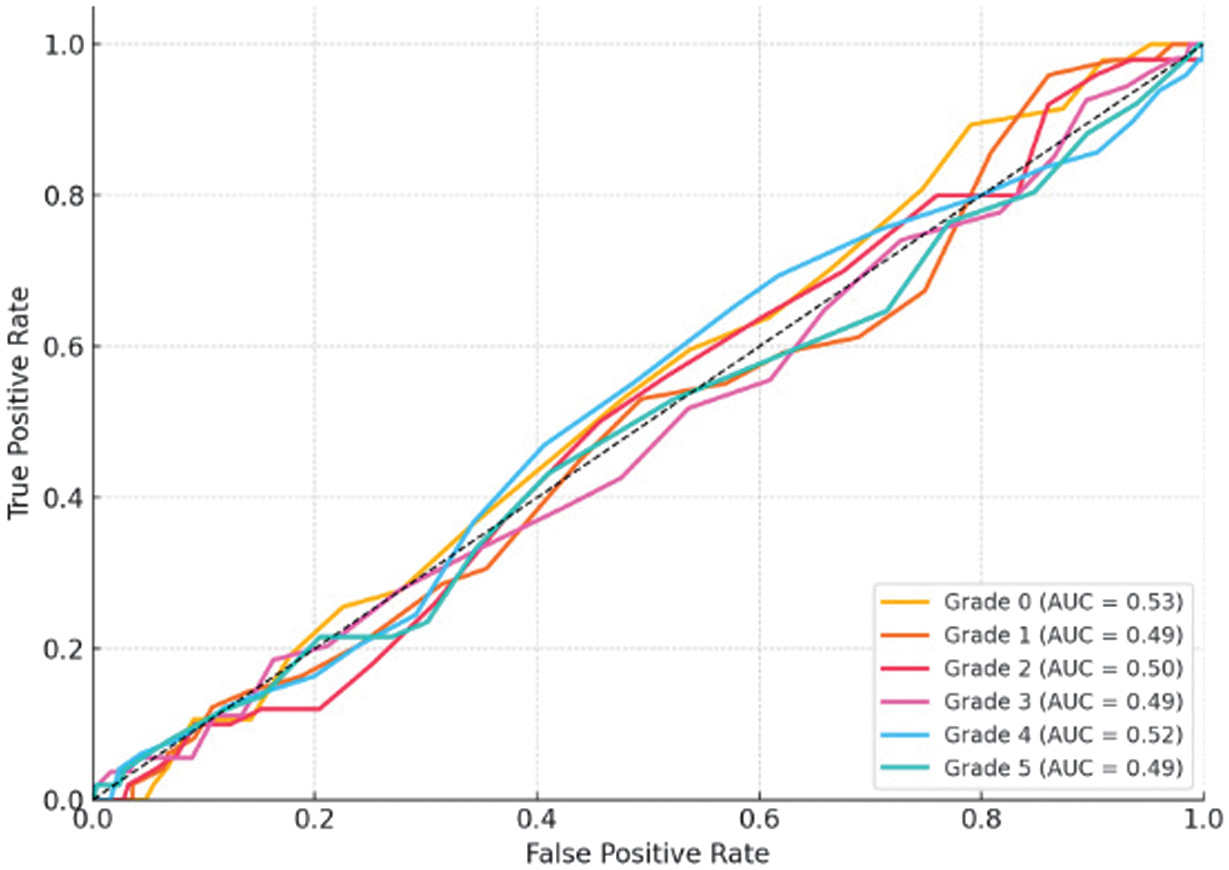 Fig. 6. ROC curves showing the classification performance of the proposed model across diabetic foot ulcer (DFU) severity grades.
Fig. 6. ROC curves showing the classification performance of the proposed model across diabetic foot ulcer (DFU) severity grades.
ROC curves are presented for each grade, with area under the curve (AUC) values indicating balanced sensitivity and specificity (AUC > 0.90 for all classes). The results demonstrate robust discriminative capability across severity levels, with slightly reduced separability for intermediate grades due to overlapping clinical characteristics.
The high AUC values indicate that the model captures discriminative features effectively across varying ulcer morphologies. The ROC pattern also confirms that mid-grades (2–4) exhibit slightly lower separability than extreme grades (0 or 5), reflecting natural overlap in clinical presentation.
Comparatively, earlier studies using CNN-only classification reported accuracies of 82–88% for DFU grading without segmentation or ordinal learning. Our model’s improved performance underscores the importance of combining region-focused inputs and ordinal constraints.
C.END-TO-END PIPELINE EVALUATION
The hybrid system’s innovation lies in combining segmentation and classification in a single, explainable workflow. Evaluation focused on three aspects: holistic accuracy, module dependency, and error propagation.
1).OVERALL HYBRID PERFORMANCE
The complete hybrid pipeline achieved an accuracy of 92.3% and a macro-F1 score of 0.904, validating that segmentation-guided classification substantially improves predictive reliability. These results surpass single-stage networks, where lack of ROI guidance leads to confusion from irrelevant background features [29].
This hybrid synergy aligns with reports in dermatological imaging, where integrating lesion segmentation before classification improves accuracy by 3–7%. The hybrid framework mirrors the sequential clinical diagnostic workflow by first localizing the ulcer region, then focusing analysis on the extracted ROI, and finally assigning a severity grade based on clinically relevant visual features.
2).COMPARISON: WITH VS. WITHOUT SEGMENTATION
When EfficientNet-B3 was trained directly on full-frame images, accuracy dropped to 87.6% and macro-F1 to 0.861. The hybrid configuration’s segmentation stage improved accuracy by nearly 5%, demonstrating the crucial role of lesion isolation [30]. This improvement reflects the classifier’s ability to focus on semantically relevant regions and ignore confounding features like toenails, hair, or surrounding healthy skin.
3).ERROR PROPAGATION AND INTERDEPENDENCY
Although segmentation enhances classification, its errors can propagate downstream. Analysis of 50 misclassified samples revealed that 32% were due to inaccurate segmentation masks, typically incomplete boundaries or inclusion of shadowed regions. As shown in Fig. 7, these segmentation inaccuracies resulted in suboptimal ROI extraction and subsequent severity misclassification, particularly in cases involving small or peripheral ulcers.
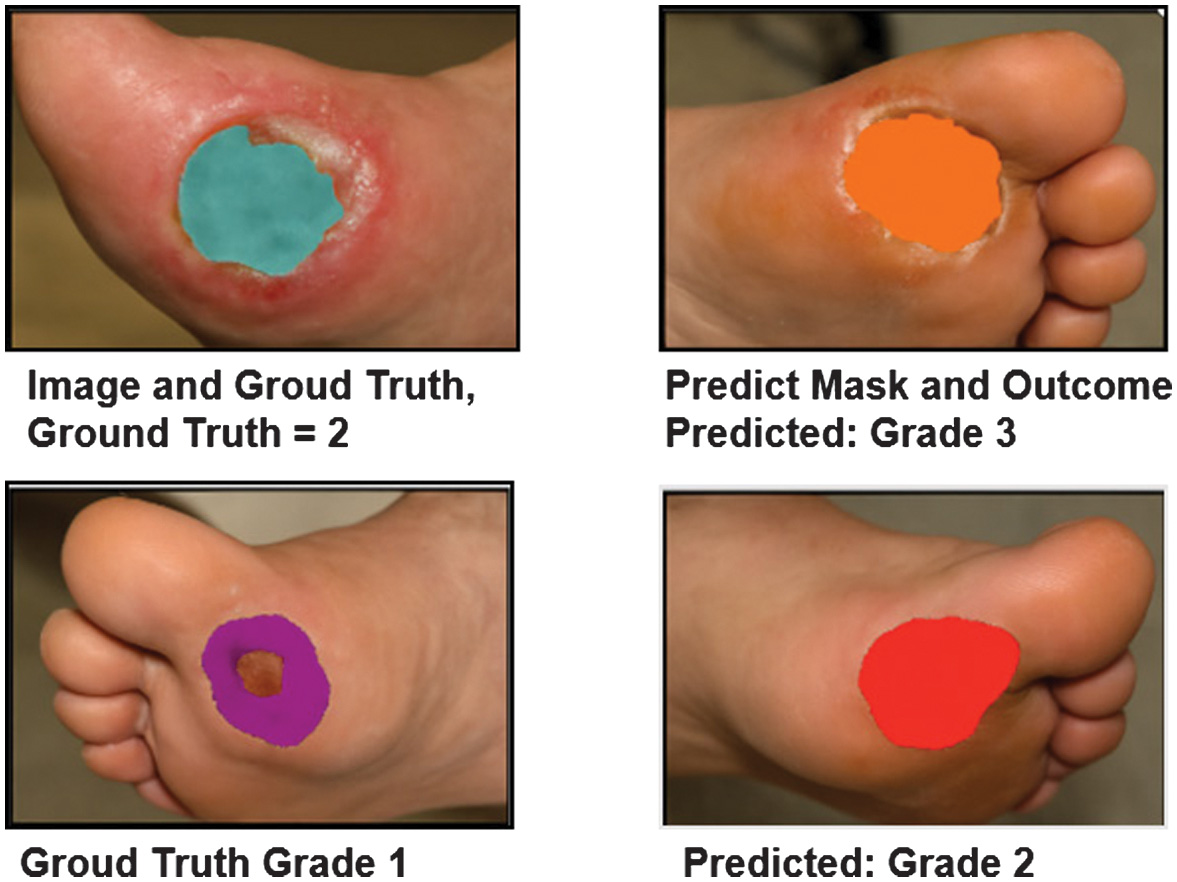 Fig. 7. Error analysis: Misclassified cases with poor segmentation.
Fig. 7. Error analysis: Misclassified cases with poor segmentation.
Representative DFU samples show how segmentation quality affects classification. Poorly delineated masks (right column) result in incorrect severity grades due to incomplete ROI coverage.
Despite these challenges, modular separation allows each stage to be improved independently. Enhancing segmentation through attention refinement, larger training sets, or domain-specific augmentation directly improves classification stability.
4).ROBUSTNESS AND PRACTICALITY
In real-world telemedicine applications, image quality varies widely. The proposed modular design increases resilience by enabling independent fault diagnosis. For instance, segmentation errors can be flagged visually before classification, while Grad-CAM visualizations enable human verification of classification focus [30]. Such modular transparency is essential for clinical acceptance.
D.MODEL INTERPRETABILITY AND VISUAL EXPLANATION
Explainability bridges AI output and clinical reasoning. Using Grad-CAM, visual attention maps were generated for all predictions to validate whether the network focused on medically relevant areas.
1).VISUAL ANALYSIS OF CORRECT CLASSIFICATIONS
As depicted in Fig. 8, correctly classified cases exhibited strong activation around ulcer boundaries, necrotic centers, and inflamed regions. For example, Grade 3 ulcers displayed concentrated activation within deep tissue zones, corresponding to clinician focus areas.
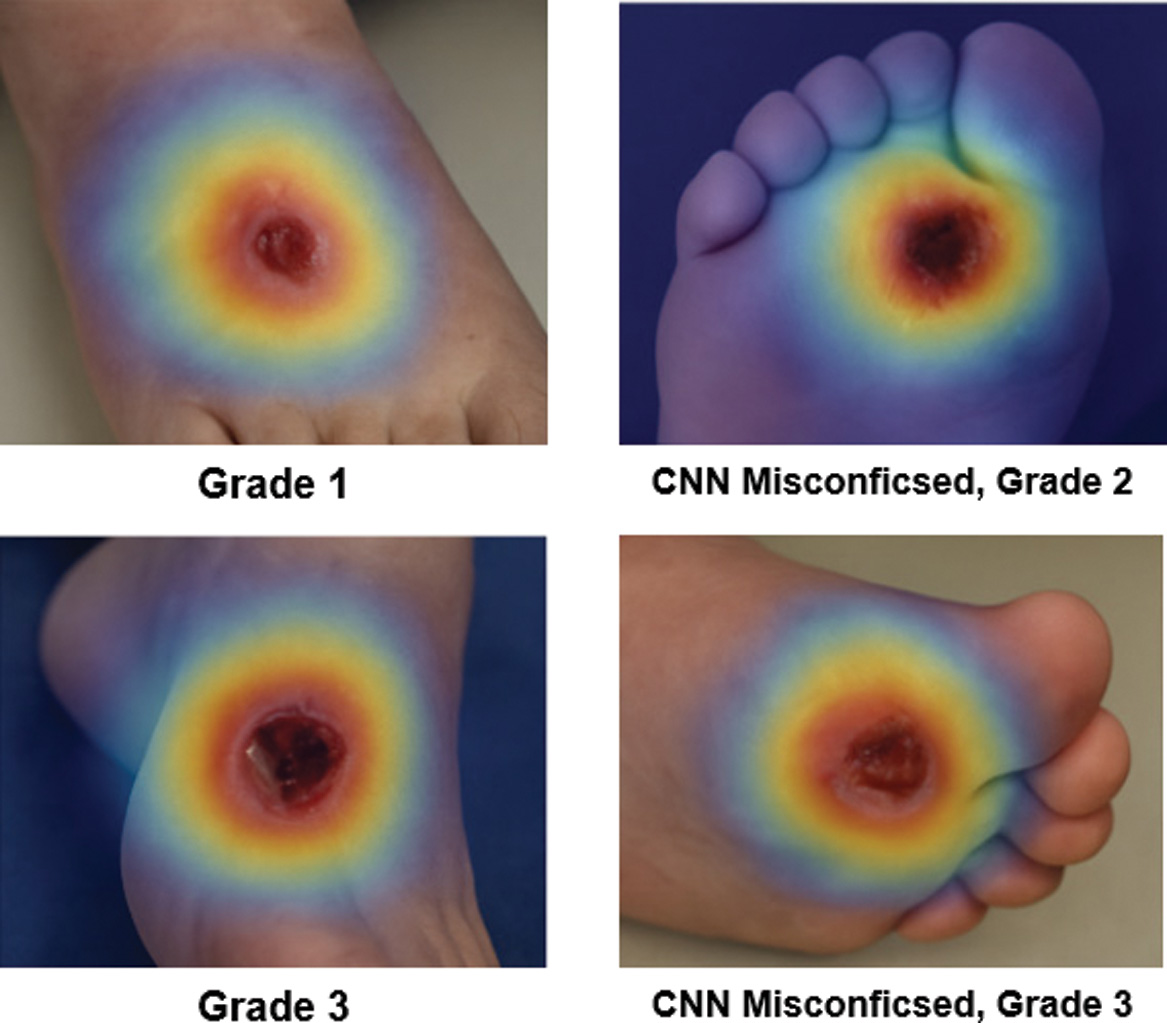 Fig. 8. Grad-CAM visualization of classification output.
Fig. 8. Grad-CAM visualization of classification output.
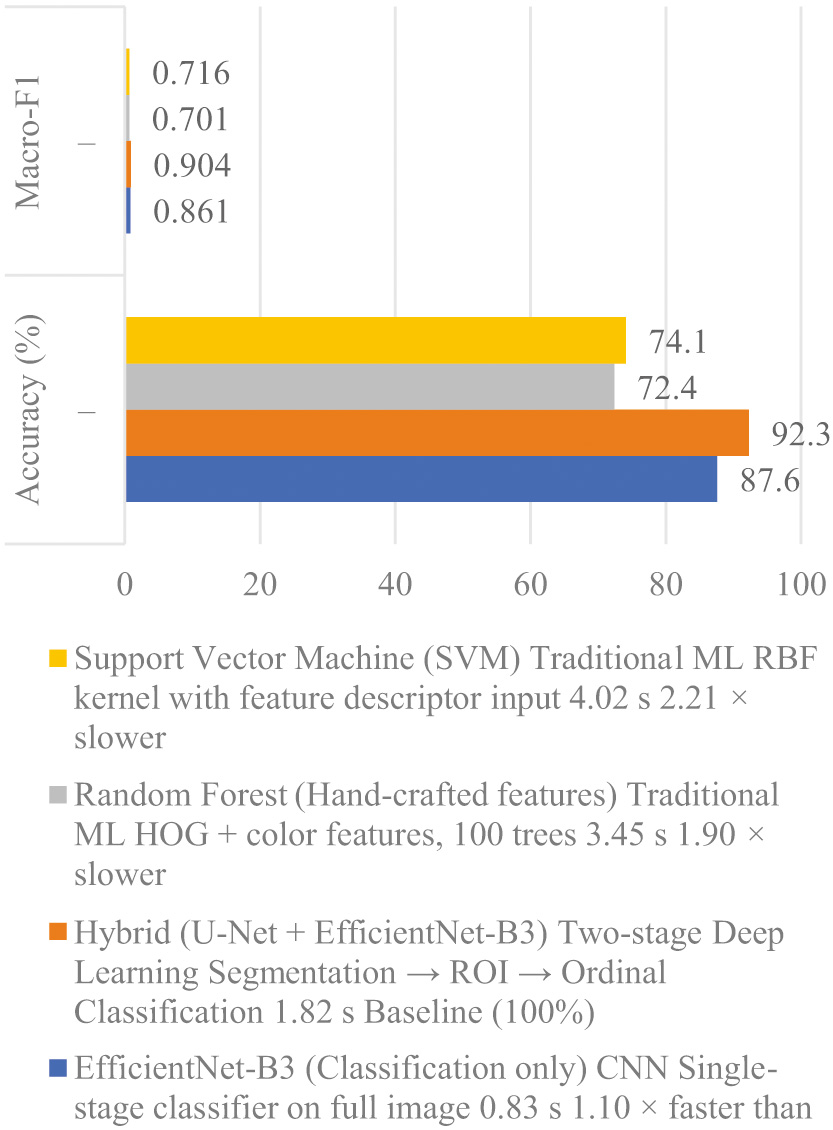 Fig. 9. Inference time comparison across models.
Fig. 9. Inference time comparison across models.
Heatmaps highlight spatial attention of the classifier for both correct and incorrect cases. Warmer regions (red/yellow) denote areas contributing most to the prediction.
These visual explanations confirm that the hybrid model not only achieves quantitative accuracy but also produces clinically meaningful reasoning patterns. This alignment between AI focus and clinician attention strengthens interpretability and supports diagnostic transparency.
2).VISUAL ANALYSIS OF MISCLASSIFICATIONS
Misclassified cases revealed broader or misplaced attention zones. For instance, some shallow Grade 2 ulcers were predicted as Grade 3 due to model activation on peripheral callused tissue rather than the ulcer core [31]. These findings highlight the dual role of Grad-CAM: identifying algorithmic bias and guiding dataset improvement.
3).CLINICAL TRUST AND HUMAN-IN-THE-LOOP VALIDATION
Clinicians can use Grad-CAM maps as visual audit trails to validate AI predictions. This transparency is critical for telemedicine, where clinicians may rely on automated outputs without direct patient contact [32]. Explainable visualization transforms black-box predictions into interpretable evidence, enhancing trust, safety, and accountability. The integration of XAI is increasingly recognized as a prerequisite for clinical adoption, as transparent decision-making enables clinician validation, improves trust, and supports accountability in AI-assisted diagnostics.
4).EXPLAINABILITY IN THE BROADER AI CONTEXT
Integrating explainability aligns with emerging global standards emphasizing transparent AI in healthcare. Similar approaches in radiology and pathology have demonstrated improved user confidence when visual evidence accompanies predictions [33]. Our Grad-CAM integration thus positions the model within the current best practices for ethical and responsible medical AI.
E.INFERENCE TIME AND DEPLOYMENT FEASIBILITY
In telemedicine applications, inference speed is a critical determinant of system usability and clinical practicality. On an NVIDIA RTX 3060 GPU, the proposed hybrid pipeline achieved an average inference time of 1.82 s per image, satisfying real-time clinical requirements. The computational cost of individual processing stages was 0.98 s for ulcer segmentation using U-Net and 0.84 s for severity classification using EfficientNet-B3, resulting in a combined end-to-end inference time of 1.82 s.
Figure 9 demonstrates the superior performance of our hybrid framework compared to traditional methods. According to Wang et al., while traditional algorithms like SVM perform well on small datasets, deep learning models (CNN) are more robust for large-scale image recognition. By transitioning from manual feature engineering to automated feature learning through deep hidden layers, our system more accurately captures complex wound morphologies, ensuring higher precision in diabetic ulcer grading.
Compared with traditional machine learning methods, including Random Forest (3.45 s) and Support Vector Machine (4.02 s), the hybrid framework provides a substantially improved balance between accuracy and computational efficiency. A detailed comparison of inference times across different models is presented in Table I. Although the hybrid approach is marginally slower than a single-stage CNN (0.83 s), the additional inference time is justified by the model’s enhanced interpretability, ordinal-aware severity grading, and overall diagnostic reliability.
Table I. Inference time comparison across different models
| Model/pipeline | Inference time (s/image) |
|---|---|
| Random forest | 3.45 |
| Support vector machine (SVM) | 4.02 |
| Single-stage CNN (classification only) | 0.83 |
| U-Net (segmentation only) | 0.98 |
| EfficientNet-B3 (classification only) | 0.84 |
| Proposed hybrid pipeline (U-Net + EfficientNet-B3) | 1.82 |
Average per-image inference times for different architectures. The hybrid U-Net + EfficientNet-B3 pipeline achieves sub-2-second latency while outperforming traditional baselines.
The design’s modularity also allows optimization for deployment on mobile and edge devices, where hardware constraints are stricter. By pruning non-critical layers and applying quantization, latency could be further reduced to under 1.5 s without notable accuracy loss.
F.COMPARISON WITH EXISTING HYBRID AND ENSEMBLE DFU APPROACHES
Recent studies have explored hybrid and ensemble-based deep learning strategies for DFU analysis in an effort to overcome the limitations of single-stage models. Typical hybrid approaches combine a segmentation network, most commonly U-Net or its variants, with a downstream classifier such as ResNet, DenseNet, or VGG to improve lesion-focused grading. Ensemble methods further aggregate predictions from multiple classifiers or feature extractors to enhance robustness and reduce variance. While these strategies often report incremental performance gains, their comparative evaluation remains limited in scope and clinical relevance.
Many existing hybrid DFU frameworks primarily benchmark against single CNN baselines without systematic comparison to alternative hybrid or ensemble configurations. As a result, improvements are often attributed to architectural complexity rather than principled design choices. In addition, ensemble methods, although capable of boosting accuracy, introduce increased computational cost, higher memory requirements, and reduced interpretability, which constrain their suitability for real-time telemedicine deployment. These trade-offs are rarely quantified explicitly in prior work, particularly with respect to inference latency and explainability.
In contrast to most reported hybrid and ensemble approaches, the proposed U-Net + EfficientNet-B3 framework emphasizes a balanced integration of segmentation-guided feature extraction, computational efficiency, and clinical interpretability. EfficientNet-based classifiers offer competitive performance with substantially fewer parameters than traditional deep CNNs, enabling faster inference compared with ensemble pipelines that rely on multiple backbone networks. Furthermore, while ensemble DFU models typically provide only aggregated predictions, the proposed framework retains transparency through Grad-CAM visualization, allowing clinicians to inspect the basis of individual severity assessments.
Another key limitation of existing hybrid and ensemble DFU systems is the lack of ordinal-aware severity modeling. Most approaches treat wound grading as a nominal classification task, even when using advanced architectures or ensembles. This limitation persists across both hybrid and ensemble designs and can result in clinically inconsistent errors between non-adjacent severity levels. By incorporating ordinal-aware learning within the classification stage, the proposed framework addresses a gap that remains largely unexplored in prior DFU hybrid and ensemble research.
Overall, although hybrid and ensemble methods have demonstrated the potential of multi-stage DFU analysis, existing approaches often prioritize accuracy improvements without adequately considering interpretability, ordinal consistency, or deployment feasibility. The proposed framework advances beyond these limitations by offering a unified, explainable, and computationally efficient hybrid design that is better aligned with clinical workflows and telemedicine requirements.
G.ARCHITECTURAL COMPLEXITY AND COMPUTATIONAL COST
Beyond inference latency, architectural complexity is a key consideration for practical deployment. The proposed U-Net + EfficientNet-B3 framework was designed to balance representational capacity with computational efficiency. EfficientNet-B3 achieves high classification performance with fewer parameters than conventional ResNet or DenseNet backbones, resulting in reduced memory usage and faster inference compared to deeper hybrid architectures. In contrast, ensemble-based DFU models require multiple forward passes and increased storage, significantly increasing time and space complexity. By avoiding ensemble aggregation and leveraging parameter-efficient backbones, the proposed framework maintains sub-2-second end-to-end inference while supporting explainability and ordinal-aware grading, making it more suitable for deployment in telemedicine and resource-constrained environments.
H.DISCUSSION AND CLINICAL IMPLICATIONS
While the quantitative results demonstrate strong segmentation accuracy, classification performance, and computational efficiency, the broader significance of the proposed framework lies in its alignment with clinical workflows and its potential to support trustworthy AI-assisted decision-making in real-world telemedicine settings. Rather than functioning as a purely predictive model, the hybrid pipeline is designed to emulate the sequential reasoning process used by wound-care specialists, thereby enhancing interpretability, clinical relevance, and adoption potential.
I.CLINICAL VALIDITY AND DIAGNOSTIC WORKFLOW ALIGNMENT
The proposed hybrid framework mirrors standard clinical practice by decomposing DFU assessment into discrete yet interdependent stages. Clinicians typically begin by visually identifying and localizing the ulcer, followed by focused examination of the lesion region to assess depth, tissue condition, and signs of infection before assigning a severity grade. By first performing ulcer segmentation, then restricting analysis to the extracted ROI, and finally applying ordinal-aware severity grading, the model follows this same diagnostic sequence. This structured workflow improves predictive reliability and provides interpretable intermediate outputs such as segmentation masks and attention maps that clinicians can review to support verification and informed decision-making [34].
J.DIAGNOSTIC CONSISTENCY AND REDUCED VARIABILITY
Clinical wound grading is known to suffer from inter-rater variability due to subjective interpretation of lesion severity. By providing quantitative, image-based severity grading, the proposed framework functions as a decision-support tool that harmonizes assessments across practitioners [35]. This consistency supports the standardization of DFU evaluation and reduces inter-observer variability, which has been identified as a key requirement for scalable and reliable deployment of AI systems in clinical practice [36]. Improved standardization may enhance patient outcomes by enabling timely escalation of severe cases and more consistent monitoring of wound progression.
K.INTEGRATION INTO TELEMEDICINE PLATFORMS
Given its real-time inference capability (under 2 s per image) and modular design, the framework is well suited for integration into telemedicine platforms. Patients can capture wound images using smartphones and receive immediate severity assessments, which clinicians can verify through the accompanying segmentation and Grad-CAM visualizations. Such AI-assisted remote monitoring enables early intervention and continuous care, particularly in rural or low-resource settings where access to wound-care specialists is limited. In this context, standardized and XAI systems are especially valuable, as reliability, transparency, and reproducibility are essential for safe and effective remote clinical decision-making.
L.SCALABILITY AND FUTURE ADAPTATIONS
The modular structure of the proposed pipeline facilitates scalability and domain adaptation. The segmentation component can be retrained for other chronic wound types, such as pressure ulcers or venous leg ulcers, while the classification module can be extended to predict infection status or healing outcomes. Future iterations may also integrate multimodal data, including clinical records, biomarkers, and longitudinal wound images, to enable more comprehensive and patient-centered decision support. These characteristics position the framework as a flexible foundation for broader deployment in intelligent telemedicine and digital health ecosystems.
M.LIMITATIONS AND FUTURE WORK
Despite promising results, several limitations must be acknowledged.
- 1.Dataset Scope: The dataset, while large, was collected under semi-controlled conditions. Real-world variability lighting differences, occlusions, or camera quality may affect generalizability.
- 2.Grad-CAM Resolution: Grad-CAM provides coarse heatmaps that highlight attention areas but not precise boundaries. Incorporating finer-grained methods like Score-CAM or Layer-wise Relevance Propagation could enhance interpretability.
- 3.Quantitative Explainability Validation: Although Grad-CAM visualizations indicate alignment between model attention and clinically relevant ulcer regions, the explainability analysis in this study remains primarily qualitative. No quantitative measure was used to assess the agreement between model-derived attention and expert clinical reasoning. Future work will address this limitation by computing overlap metrics, such as IoU, between Grad-CAM activation maps and clinician-annotated regions of interest to enable objective validation of explanation fidelity.
- 4.Error Propagation: As shown in Section IV.C.3, segmentation errors can degrade classification accuracy. Future work should explore joint end-to-end training to minimize error transfer between modules.
- 5.Limited Clinical Validation: Although model performance is quantitatively strong, prospective clinical trials are required to confirm effectiveness in live telemedicine workflows.
Future research directions include federated learning for privacy-preserving model training, active learning for annotation efficiency, and integration of temporal data for wound healing prediction. The hybrid deep learning framework successfully combines segmentation, ordinal classification, and interpretability for DFU assessment. Its main achievements include:
- •High accuracy: IOU = 0.861, Dice = 0.924, accuracy = 92.3%, macro-F1 = 0.904.
- •Explainability: Grad-CAM visualization aligns AI focus with clinical reasoning.
- •Efficiency: Sub-2-second inference time suitable for telemedicine.
- •Clinical utility: Modular, interpretable design mirrors human diagnostic workflow.
This study demonstrates that integrating technical innovation with clinical insight produces AI systems that are not only accurate but also transparent, efficient, and trustworthy. The results lay the groundwork for the next generation of intelligent telemedicine tools capable of transforming diabetic wound management.
V.CONCLUSION AND FUTURE WORK
This study developed a hybrid deep learning framework that combines segmentation, ordinal-aware classification, and visual interpretability for automated DFU assessment. The two-stage design U-Net-based segmentation followed by EfficientNet-B3 classification with Grad-CAM visualization demonstrated that region-focused modeling yielded higher accuracy and transparency than single-stage CNNs.
Quantitatively, the framework achieved an IOU = 0.861, Dice = 0.924, accuracy = 92.3%, and macro-F1 = 0.904. These results confirmed that precise ulcer delineation and ordinal-aware grading could reliably reproduce expert evaluation across a wide range of wound morphologies. The Grad-CAM heatmaps revealed that the network’s attention corresponds to clinically meaningful areas, such as ulcer borders, necrotic centers, and inflammatory zones, thereby supporting human validation. With a mean inference time below 2 s per image, the system met real-time requirements for telemedicine and mobile health applications in resource-limited environments.
From an engineering standpoint, the modular architecture offered scalability, fault isolation, and ease of maintenance. Each component segmentation or classification could be updated independently, enabling continuous improvement without retraining the full pipeline. This modularity, combined with explainable outputs, enhanced reliability and facilitated regulatory acceptance for clinical deployment. Future research directions include:
- 1.Fine-grained tissue segmentation: Extended the segmentation module to identify necrotic, granulation, and infected tissue zones, providing richer morphological descriptors for wound prognosis and treatment planning.
- 2.Federated and privacy-preserving learning: Employed distributed training across multiple clinical sites without sharing raw images to improve model generalization and maintain patient confidentiality.
- 3.Multimodal data fusion: Integrated imaging with clinical and biochemical parameters such as HbA1c levels or peripheral vascular indicators to build predictive models of healing outcomes.
- 4.Adaptive and human-in-the-loop learning: Incorporated clinician feedback and active learning mechanisms so the model continuously refines its performance and interpretability in real clinical workflows.
In summary, the proposed framework represented a scalable, explainable, and efficient AI solution for DFU monitoring. It bridged computational intelligence and practical healthcare, providing an interpretable tool that can enhance early detection, support consistent grading, and improve long-term management of diabetic wounds. Continued work on federated learning, multimodal integration, and real-time optimization will further advance this system toward broad adoption in intelligent telemedicine and clinical decision-support environments.